- Title
-
REREa/Atrophin-2 interacts with histone deacetylase and Fgf8 signaling to regulate multiple processes of zebrafish development
- Authors
- Plaster, N., Sonntag, C., Schilling, T.F., and Hammerschmidt, M.
- Source
- Full text @ Dev. Dyn.
|
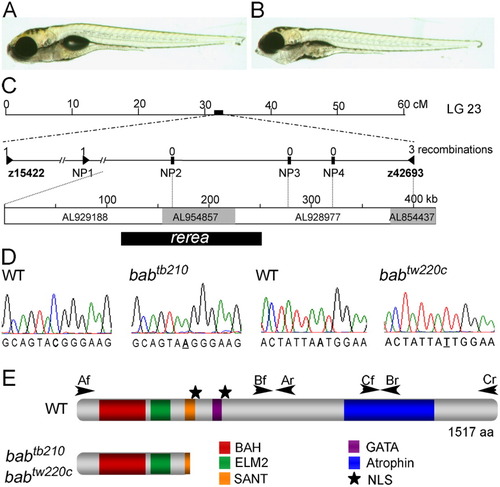
The bab phenotype is caused by a disruption in rerea. A,B: The wild-type (WT, A) and bab (B) larvae at 5 days postfertilization (dpf) are shown in lateral view. Ventral distension of the pharyngeal area in bab is caused by the inverted second arch cartilage. C: The bab locus is at approximately 32 cM on linkage group 23. Expanded view of the genetic map shows the number of recombinations in 2,094 alleles above the marker name. The genomic contigs that span the completely linked region, their Genbank accession numbers, and the position of rerea are shown below. D: Sequence chromatographs of both the babtb210 and babtw220c in comparison with wild-type sequence are shown with the mutation underlined. E: The REREa schematic displays the positions of identified protein domains with colored boxes and nuclear localization signals (NLS) with stars. Arrowheads mark the location and direction of primers used to amplify cDNA. Nonsense mutations in both alleles of bab truncate REREa in the SANT domain. PHENOTYPE:
|
|
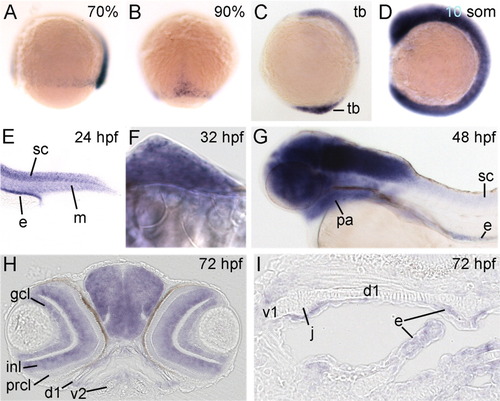
rerea is expressed dynamically in all developmental stages examined. All panels show whole-mount in situ hybridizations of rerea for wild-type embryos or larvae of ages indicated in upper right corner. A-G: Lateral views with anterior to the left (A,C,D,G), dorsal view with animal pole up (B), lateral view on tail (E), and frontal view of fin bud with medial to the left (F). H,I: Frontal section though head (H) and horizontal sections with anterior to the left through the pharyngeal arches (I). d1, dorsal cartilages of arch 1; e, endoderm; gcl, ganglion cell layer; inl, inner nuclear layer; j, joint; m, somitic mesoderm; pa, pharyngeal arches; prcl, photoreceptor cell layer; sc, spinal cord; tb, tail bud; v1 and v2, ventral cartilages of arches 1 and 2, respectively. EXPRESSION / LABELING:
|
|
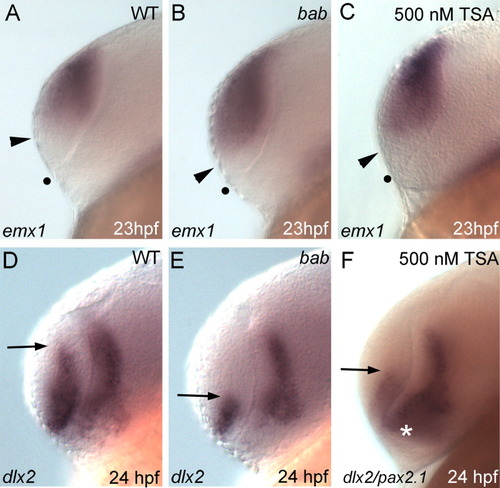
Disruption of REREa or histone deacetylase (HDAC) activity causes a dorsalization of the telencephalon. A-F: All panels show lateral views of the telencephalon in whole-mount in situ hybridizations on wild-type (WT, A,D), bab (B,E) or Trichostatin A (TSA) -treated (C,F) embryos. Probes are denoted in the lower left corner with stages in the lower right. A-C: emx1 expression extends ventrally from the dorsal telencephalon to the arrowhead and is expanded in bab and TSA-treated embryos. The dot marks the ventral telencephalic boundary. D-F: This results in a reduction of dlx2 expression in the telencephalon. Arrows mark the dorsal limit of dlx2 expression. pax2.1 expression (white asterisk) in F overlaps the diencephalic expression of dlx2, but is not expressed in the telencephalon. EXPRESSION / LABELING:
PHENOTYPE:
|
|
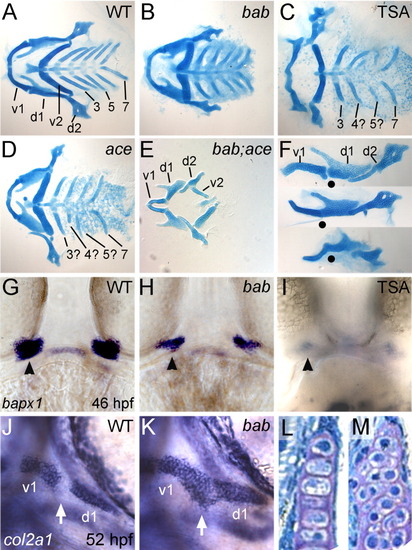
rerea genetically interacts with fgf8 in pharyngeal arch development, and disruption of histone deacetylase (HDAC) activity results in bab-like fusions of the first arch. A-M: Alcian blue-stained cartilage flat-mounts at 5 dpf (A-F), whole-mount in situ hybridizations of embryos with bapx1 (G-I) and col2a1 (J,K), and frontal sections through the palatoquadrate (d1 element, L,M) are shown for wild-type (WT; A,F top, G,J,L), bab (B,F middle, H,K,M), Trichostatin A (TSA) -treated (C,F bottom, I), ace (D), and bab;ace (E) embryos or larvae. Anterior is to the left in all images except G-I,L,M, which are frontal views on the mouth. A-E: Ventral views of flat-mounted cartilages with posterior arches numbered 3-7, reveal the presence but reduction in all elements in bab (B), a missing posterior arch in both TSA-treated (C) and ace (D), and a complete loss of posterior arches in bab;ace (E). Disruption of REREa (H) or TSA treatment (I) results in reduction of joint specification marker, bapx1 compared with wild-type embryos (G). Later col2a1 is misexpressed in cells of the presumptive joint (white arrow) in bab (K) compared with WT (J) leading to a fusion of the first arch cartilages (F, lateral view of arches 1 and 2) in wild-type (top F), bab (middle F), and TSA-treated (bottom F) embryos. Dots mark the position of the joint in each preparation. Chondrocyte morphology at 4 hpf is round in bab (M) compared with the stacked structure in wild-types (L). d1 and d2, dorsal cartilages of arches 1 and 2, respectively; v1 and v2, ventral cartilages of arches 1 and 2, respectively. |
|
rerea interacts genetically with fgf8 in posterior mesoderm development. A-C: Lateral views of live 24 hpf embryos with genotypes in the upper right corner. A′-C′: Inset panels show the corresponding lateral views of diphosphorylated ERK1&2 (pERK) immunostained tail buds at 13 somites. Arrows denote the position of pERK staining. D-I: Whole-mount in situ hybridizations of papc at 4 somites (D-F) or myod with Ntl immunostains at 16 somites (G-I) shown in dorsal view with genotypes shown in the upper right corner. The arrowhead in I marks a gap in Ntl staining. "WT(bab)" notes that there is no observable difference between wild-type (WT) and bab embryos. ace shows a slight reduction in the amount of posterior mesoderm (B) and presomitic mesoderm as seen by papc expression at four somites (E), while bab;ace (C,F) shows a severe reduction in comparison to wild-type or bab (A,D). EXPRESSION / LABELING:
PHENOTYPE:
|
|
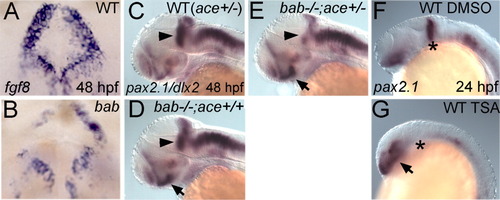
rerea and fgf8 genetically interact in the midbrain-hindbrain boundary (MHB), and disruption of REREa or inhibition of histone deacetylase (HDAC) activity results in the inability to maintain the MHB. A-F: All panels show whole-mount in situ hybridizations in frontal view (A,B) or lateral view (C-G) of 48 hours postfertilization (hpf; A-E) or 24 hpf (F,G) embryos. Genotypes are shown in the upper right corner, whereas probes and stages are shown in the lower left and right corners, respectively. A,B: bab (B) displays degeneration of the MHB as seen by the patchy disappearance of fgf8-expressing cells compared with WT embryos (A). C,D: pax2.1 is expressed in the MHB (arrowheads) in wild-type (WT; ace+/-, C) and bab-/-;ace+/+ (D) embryos at 48 hpf. E: Reduction of Fgf8 in bab-/-;ace+/- leads to a reduction of pax2.1 in the MHB. dlx2 expression was used for the purpose of identifying bab embryos and does not interfere with the MHB staining of pax2.1. F,G: The 350 nM Trichostatin A (TSA) treatment from dome stage to 24 hpf also leads to a loss of pax2.1 expression at the MHB (asterisk in G) compared with wild-type embryos treated with dimethylsulfoxide (DMSO) alone (F). C-G: pax2.1 expression in the eye (arrows) is up-regulated in bab (D,E) and TSA-treated embryos (G) compared with wild-types (C,F). |
|
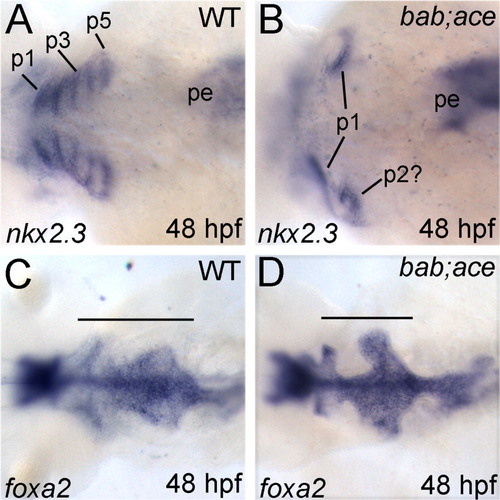
rerea and fgf8 interact during formation of the endodermal pouches. A-D: All panels show dorsal views, with anterior to the left, of whole-mount in situ hybridizations of wild-type (A,C) and bab;ace (B,D) embryos. Probes are shown in the lower left corner, whereas stages are in the lower right. Combined disruption of rerea and fgf8 leads to loss of the posterior endodermal pouches as seen by nkx2.3 expression (A,B) while medial endoderm (foxa2) remains (C,D). Horizontal bars (C,D) mark the anterior-posterior extent of the pharyngeal endoderm. p1-5, endodermal pouches 1-5; pe, posterior endoderm. |
|
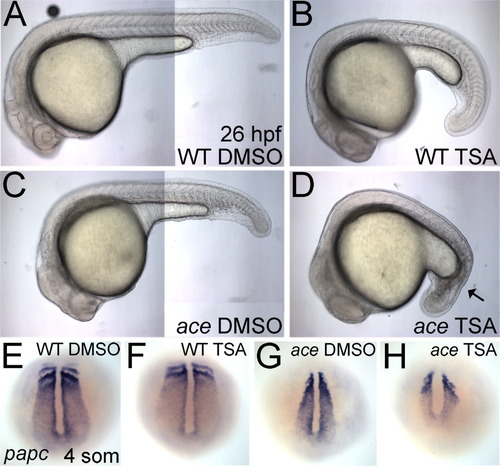
Histone deacetylase (HDAC) activity is required for proper Fgf signaling during development of the posterior mesoderm. A-D: Lateral views of live 26 hours postfertilization (hpf) wild-type (A,B) or ace (C,D) embryos, treated with either dimethylsulfoxide (DMSO) control medium (A,C) or 200 nM Trichostatin A (TSA, B,D) are shown. TSA-treated embryos are slightly delayed. The black arrow in D marks severely disrupted tail mesoderm, resembling the bab;ace phenotype at 24 hpf. E-H: Whole-mount in situ hybridizations (dorsal views on the tail bud) of papc at the four-somite stage in wild-type embryos (E,F) or ace (G,H) treated with DMSO control medium (E,G) or 200 nM TSA (F,H) from 30% epiboly. EXPRESSION / LABELING:
PHENOTYPE:
|
|
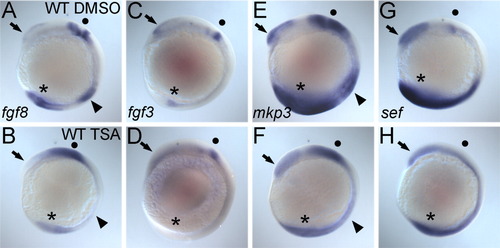
The proper expression of fgf3 and fgf8 and fgf target genes, sef and mkp3, requires histone deacetylase (HDAC) activity. A-H: All panels show lateral views of whole-mount in situ hybridizations of seven-somite stage WT embryos treated with dimethylsulfoxide (DMSO, A,C,E,G) or 350 nM Trichostatin A (TSA, B,D,F,H) from dome to the seven-somite stage. Probes are shown in the bottom left corner of each wild-type panel. A,B,E,F: The forebrain (arrows) and tail bud (asterisk) expression of all markers is decreased or absent in TSA-treated embryos while the midbrain-hindbrain boundary (MHB) domains (dots) are decreased and diffuse. Both fgf8 (A,B) and mkp3 (E,F) expression in the developing somites (arrowheads) is also lost in TSA-treated embryos. EXPRESSION / LABELING:
|
|
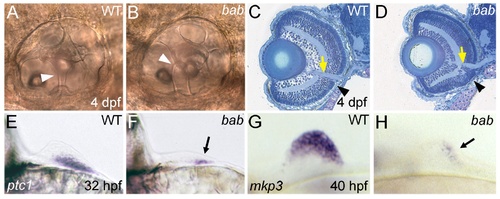
REREa is necessary for the proper development of multiple tissue types. (A,B) Lateral views with anterior to the left of otic capsules (ears) in live larvae at 4 dpf. The epithelial pillars (white arrowheads) that help to form the semicircular canals of the otic capsule fail to fuse in bab (B) compared to wild-type (A) larvae. (C,D) Methylene blue/azure II stains of frontal plastic sections through the head of wild-type (C) and bab (D) 4 dpf larvae. The yellow arrows mark where retinal tissue has merged with the optic nerve, while the black arrowheads mark disrupted retinal lamination in the ventral retina of bab compared to wild-type larvae. Note the compacted appearance of the bab retina compared to wild-type. (E-H) Views on the fin buds of whole-mount in situ hybridizations of wild-type (E,G) and bab (F,H). (E,F) Frontal view with medial to the left of ptc1 expression at 32 hpf. (G,H) Lateral view with anterior to the left of mkp3 expression at 40 hpf. Arrows in (F) and (H) denote the residual fin bud expression of ptc1 and mkp3, respectively. In both cases (ptc1 and mkp3), expression of these genes is initiated but not maintained in the fin bud. |
|
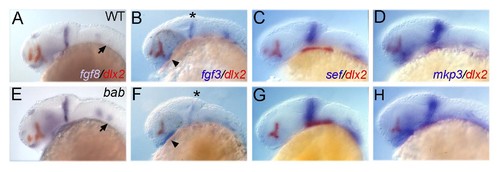
bab displays tissue-specific up- and down-regulation of Fgf pathway transcripts at 24 hpf. All panels show lateral views of whole-mount in situ hybridizations of 24 hpf WT (A-D) or bab (E-H) embryos for fgf8 (A,E), fgf3 (B,F), sef (C,G), and mkp3 (D,H) in blue and dlx2 in red (for genotyping purposes). Expression in the diencephalon (arrowheads), posterior otic capsule (arrows), and MHB (*) is denoted for the corresponding transcripts. EXPRESSION / LABELING:
|

Unillustrated author statements EXPRESSION / LABELING:
|