Fig. 4
|
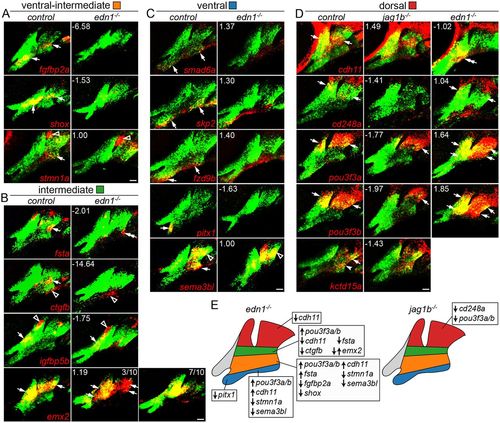
Changes in domain-specific gene expression in edn1 and jag1b mutants. Two-color fluorescent in situ hybridizations were performed for genes of interest (red) and dlx2a (green) to label arch CNCCs at 36 hpf. Numbers indicate the gene expression fold-change in mutant versus wild-type fli1a:GFP+; sox10:dsRed+ CNCCs as determined by RNAseq. Maximum intensity projections show the mandibular (left) and hyoid (right) arches. (A) The ventral-intermediate expression of fgfbp2a, shox and stmn1a is lost in edn1 mutants, yet pouch expression of stmn1a is unaffected. (B) In edn1 mutants, intermediate expression of fsta and ctgfb is lost, emx2 is variably upregulated or downregulated, and CNCC and mesoderm expression of igfbp5b is unaltered. We also note some ectopic ventral hyoid fsta expression in edn1 mutants. (C) Ventral expression of smad6a, skp2 and fzd9b is normal in edn1 mutants, yet ventral expression of pitx1 and sema3bl is lost. Note that dorsal expression of sema3bl is unaffected. (D) In jag1b mutants, dorsal expression of cd248a is lost, pou3f3a and pou3f3b are reduced, and cdh11 and kctd15a are unaffected. In edn1 mutants, cdh11 expression shifts to ventral CNCCs, pou3f3a and pou3f3b are ectopically expressed in ventral CNCCs, and cd248a is largely unaffected. Arrows indicate expression in predicted arch domains, open arrowheads indicate additional expression domains, and double arrows show expansion into other CNCC domains in mutants. Unless stated otherwise, consistent expression patterns were seen in a minimum of three wild types and three mutants for each experiment. Scale bars: 20 μm. (E) Summary of verified gene expression changes in edn1 and jag1b mutants. Unaffected genes are not listed. |
| Fish: | |
|---|---|
| Observed In: | |
| Stage: | Prim-25 |