- Title
-
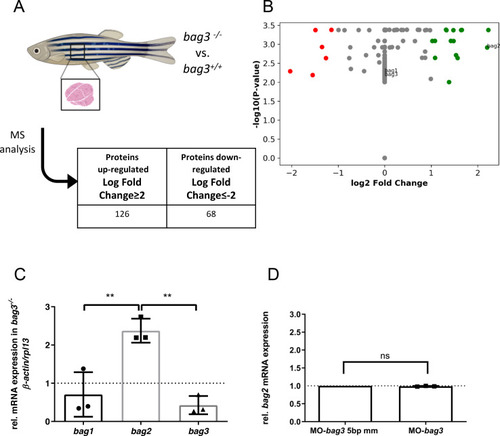
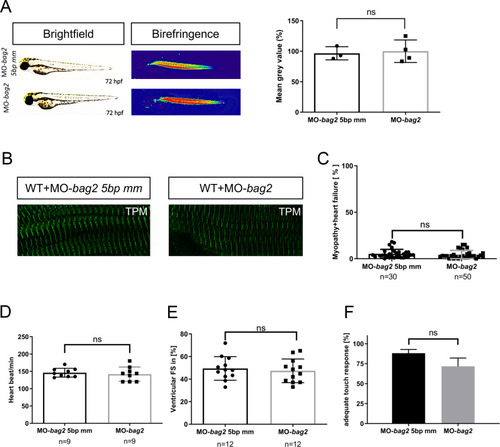
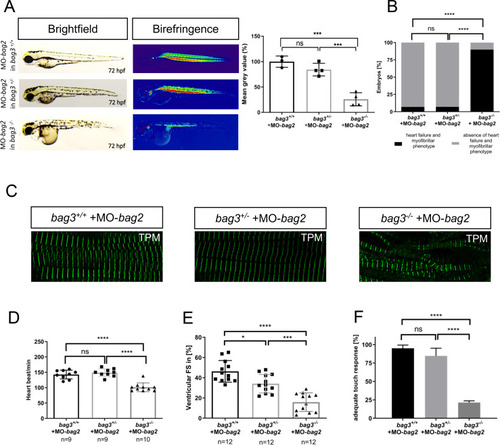
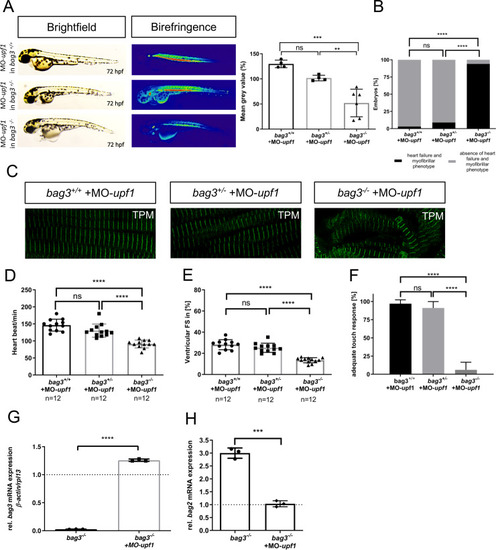
Genetic compensation prevents myopathy and heart failure in an in vivo model of Bag3 deficiency
- Authors
- Diofano, F., Weinmann, K., Schneider, I., Thiessen, K.D., Rottbauer, W., Just, S.
- Source
- Full text @ PLoS Genet.
|
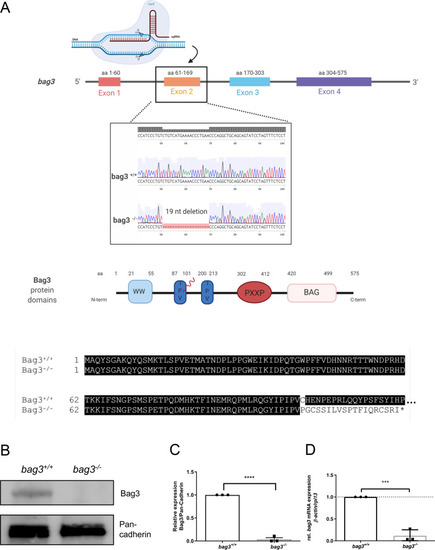
EXPRESSION / LABELING:
PHENOTYPE:
|
|
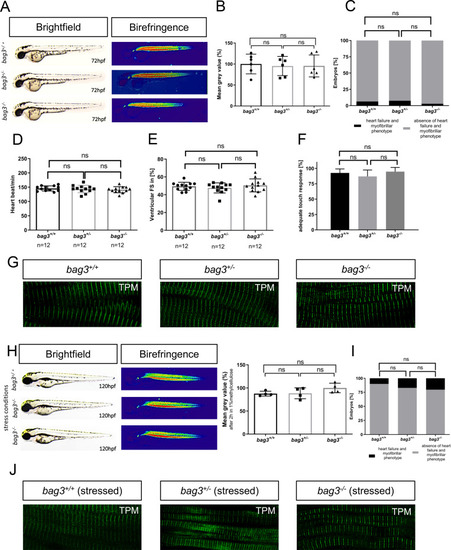
PHENOTYPE:
|
|
PHENOTYPE:
|
|
PHENOTYPE:
|
|
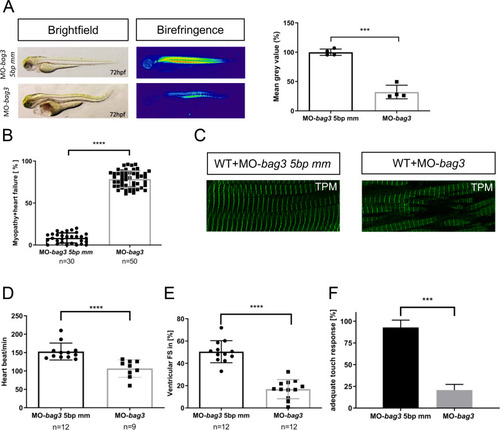
EXPRESSION / LABELING:
PHENOTYPE:
|
|
PHENOTYPE:
|
|
PHENOTYPE:
|
|
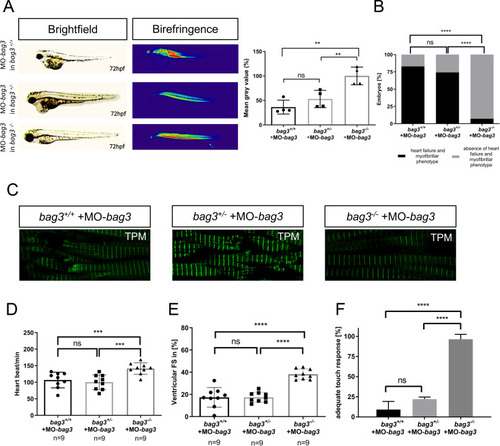
EXPRESSION / LABELING:
PHENOTYPE:
|