- Title
-
Tissue specific roles for the ribosome biogenesis factor Wdr43 in zebrafish development
- Authors
- Zhao, C., Andreeva, V., Gibert, Y., LaBonty, M., Lattanzi, V., Prabhudesai, S., Zhou, Y., Zon, L., McCann, K.L., Baserga, S., and Yelick, P.C.
- Source
- Full text @ PLoS Genet.
|
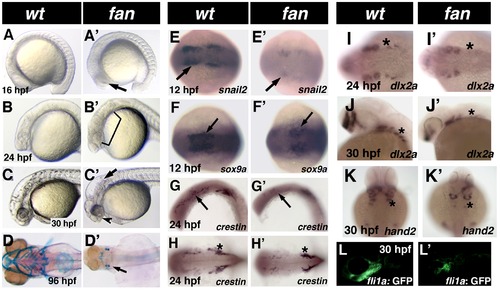
Phenotype of fan mutants. (A–C2) Live images of developmentally staged wild type (A–C) and fan mutant (A2–C2) zebrafish. Arrow in A2 indicates necrotic cells in the presumptive eye region. Bracket in B2 shows necrosis in neural and pharyngeal arch tissues. Arrow in C2 points to the distinct hydrocephaly in fan mutant hind brain ventricles, arrowhead indicates incomplete choroid fissure closure and craniofacial defects. (D–D2) Alcian blue stained pharyngeal arch cartilages in 4dpf wild type (D) and fan mutant (D2). (E–K2) Whole mount ISH images of wild type (E–K) and fan mutants (E2–K2) for neural crest markers at indicated developmental stage. Arrows in (E2–G2) and asterisks in (I2–K2) indicate reduced gene expression in fan mutant embryos. Interestingly, fan mutant embryos exhibit similar crestin expression in the trunk NCCs (asterisks in H, H2) but reduced expression in cranial NCCs. (L–L2) Tg(fli1a:EGFP)/fan mutants (L2) exhibit reduced GFP expression in the pharyngeal arch region as compared to wild type embryos (L). EXPRESSION / LABELING:
PHENOTYPE:
|
|
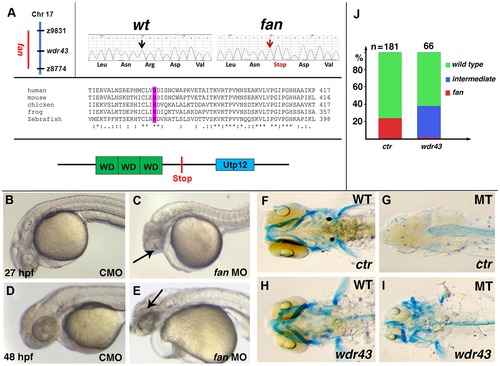
fan encodes zebrafish Wdr43/Utp5. (A) Left side: Chromosomal position of the fan locus (left). Sequencing trace data of wild type and fan mutant alleles, with red arrow indicating a premature stop codon in fan mutants (right). Comparison of the conserved nature of the amino acid mutated in fan zebrafish (purple) (middle). Schematic of the Wdr43 protein domains and fan mutant predicted premature stop codon (bottom). (B–E) Live images of zebrafish treated with control MO (CMO) or fan MO at indicated developmental times. (F–I) Alcian blue stained wild type (F, H) and fan mutant (G, I) embryos injected at the single cell stage with GFP control (ctr) or wild type wdr43 mRNA. (I) Injected wdr43 mRNA rescued fan mutant cartilage formation. (J) Percentage of control and wdr43 mRNA injected fan mutant clutches exhibiting wild type, intermediate and fan mutant pharyngeal arch cartilage formation. Numbers of injected embryos scored are indicated at the top of each bar. |
|
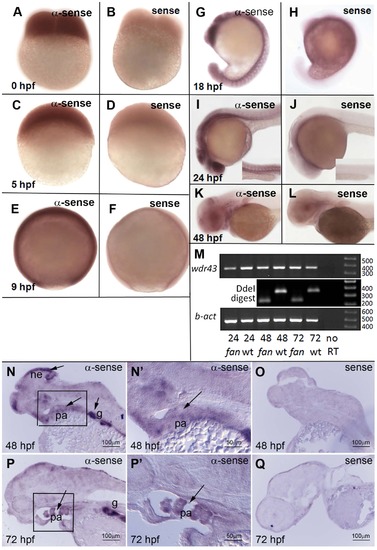
Developmental expression pattern of zebrafish wdr43 mRNA. (A–L) Whole mount ISH images of sense and anti-sense wdr43 probes in developmentally staged zebrafish embryos. (M) RT-PCR results of wdr43 gene expression in wild type and fan mutant embryos at indicated times (hpf). fan mutant mRNA was detected at 48 and 72 hpf, as indicated by digestion with DdeI, a unique restriction site generated by the fan point mutation. (N–Q) Sectioned ISH revealed discrete wdr43 mRNA expression in neurepithelium (ne), pharyngeal arch tissues (pa), and gut (g) (arrows). (N2, P2) Higher magnification images of boxed regions in N, P, respectively. Sense controls did not exhibit staining (O, Q). EXPRESSION / LABELING:
|
|
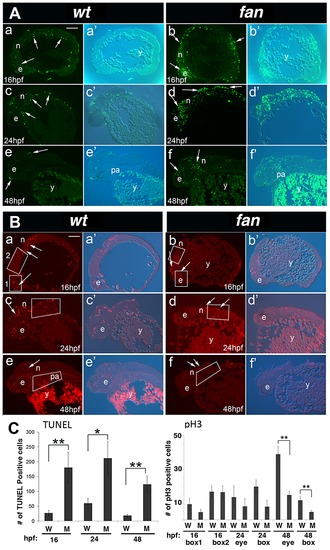
fan mutants exhibit increased apoptosis and reduced cell proliferation. (A) TUNEL Staining. Wild type embryos at 16 hpf (a, a2), 24 hpf (c, c2) and 48 hpf (e, e2) exhibited few apoptotic cells at all stages (arrows). In contrast, age matched fan mutant embryos exhibited increased levels of apoptotic cells at all stages examined (b, b2, d, d2, f, f2, arrows). Abbreviations: e, eye; n, neural tissue; pa, pharyngeal arches; y, yolk). Scale bar = 100 µm. (B) pH3 Immunofluorescent (IF) histochemistry. At 16 hpf, 24 hpf and 48 hpf, wild type embryos exhibit discrete pH3 expression in proliferating cells of the eye (e), neural tissues (n), and pharyngeal arches (pa) (a, a2, c, c2, e, e2). In contrast, age matched fan mutants exhibited reduced pH3 positive cell proliferation at all stages examined (b, b2, d, d2, f, f2, arrows). (Abbreviations: y, yolk). Scale bar = 100 µm. For both (A) and (B), fluorescent (a–f) and bright field plus fluorescent (a2–f2) images are shown. (C) Quantification of TUNEL and pH3 IF. For TUNEL, all apoptotic cells in each panel were counted for comparison between age matched wild type and fan mutant embryos. For pH3 IF, red fluorescent cells were counted in boxed areas as indicated. These results showed that fan mutants exhibited significantly increased apoptosis at all stages examined and significantly reduced cell proliferation at 48 hpf. At least 3 sectioned embryos were examined for each genotype at each developmental stage. Statistical analyses were performed using Student′s t-test. (p = >0.01). PHENOTYPE:
|
|
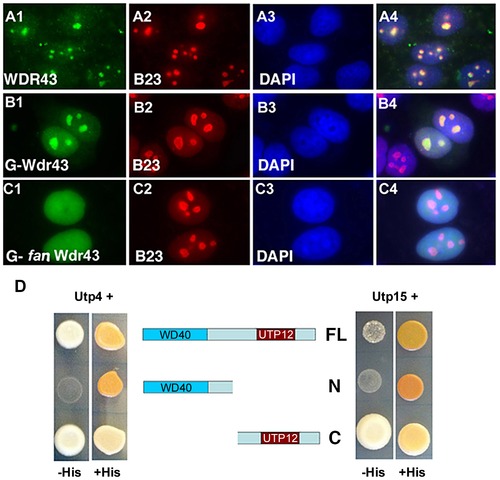
Subcellular localization and Y2H analyses of wild type and fan mutant Wdr43. (A1–A4) IF images of HeLa cells immunostained with anti-WDR43 (green) and anti-B23 (red) antibodies followed by DAPI stain (blue) to visualize nuclear DNA. IF images of Hela cells transfected with EGFP tagged zebrafish wild type Wdr43 (B1–B4) or fan mutant Wdr43 (C1–C4). Anti-GFP antibody was used to increase the fluorescent signal of EGFP-tagged wild type and fan mutant Wdr43 expressed proteins. Stained cells were counterstained with anti-B23 (red) and DAPI (blue). (D) Y2H analysis of zebrafish full length (FL) Wdr43, truncated fan mutant Wdr43 (N), and Wdr43 C-terminal domain (C) interactions with zebrafish Utp4 and Utp15 proteins. |
|
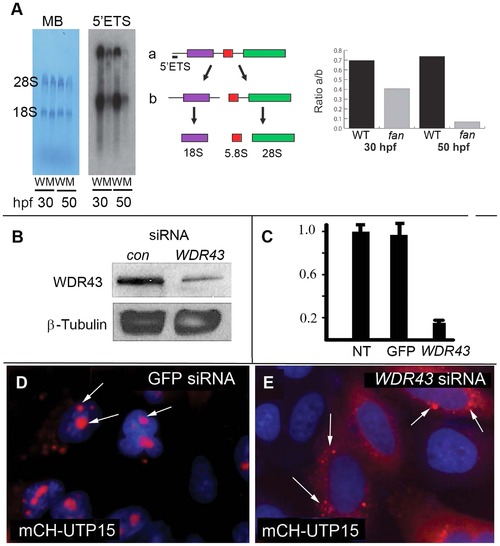
Ribosome biogenesis defects in fan mutant zebrafish. (A) Northern blot analysis of rRNA isolated from 30 and 50 hpf wild type (W) and fan mutants (M) using an oligonucleotide probe against the 52ETS region of zebrafish pre-rRNA. Pre-rRNAs (a) and (b) are indicated. Methylene blue (MB) staining of the mature 28S and 18S rRNAs was carried out as a loading control. Quantitation of the ration of a/b was performed using Image J. (B) Western blot of whole-cell extracts treated with WDR43 or EGFP control siRNA as indicated. (C) qRT-PCR analyses of human WDR43 gene expression level in non-treated (NT), GFP or WDR43 siRNA treated HeLa cells normalized to β -actin. (D, E) Subcellular localization of mCherry-tagged Utp15 (red) in control GFP (D) and WDR43 (E) siRNA treated HeLa cells counter stained with DAPI (blue). |
|
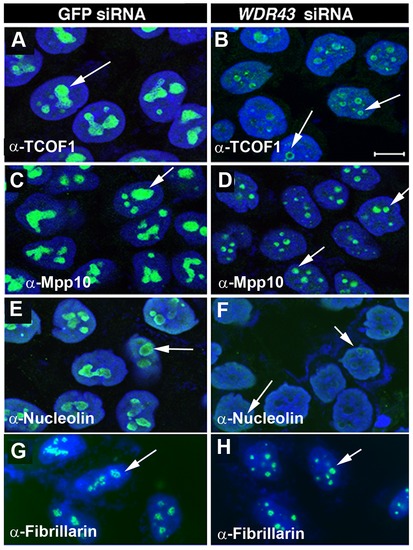
Subcellular localization of nucleolar proteins in WDR43 depleted HeLa cells. IF analysis of nucleolar protein localization in control siRNA (A, C, E, G) and WDR43 (B, D, F, H) siRNA treated HeLa cells. WDR43 depleted cells exhibited mislocalized expression of TCOF1 (B), Mpp10 (D), Nucleolin (F) and Fibrillarin (H) (arrows), as compared to their respective control GFP siRNA treated cells (see arrows). Scale bar = 10 μm. |
|
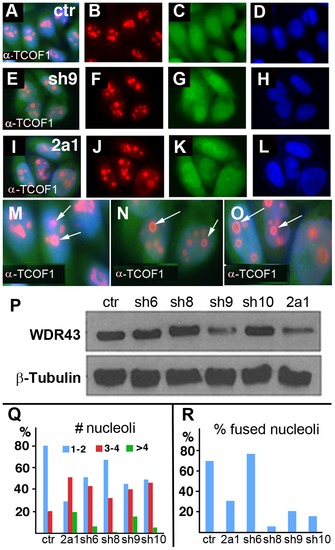
TCOF1 expression in stable WDR43 shRNA cell lines. (A–O) IF analysis of TCOF1 (red) localization in control shRNA (A–D, M) and human WDR43 shRNA (E–L, N, O) stable HeLa cell lines. All cells were positive for shRNA expression (green) and DAPI stained nuclei (blue). (P) Western blot analyses of WDR43 expression in stable WDR43 shRNA cell lines. (Q–R) Quantification of TCOF1 positive nucleoli per cell (Q), and percent cells with fused TCOF1 positive nucleoli (R), in control and WDR43 shRNA stable lines. The number of nucleoli was increased (Q), and the percentage of fused nucleoli was reduced (R) in WDR43 depleted cells, as compared to control cells. Sample numbers: Panel Q - ctr 290, 2a1 136, sh6 195, sh8 172, sh9 172, sh10 108; Panel R – ctr 268, 2a1 115, sh6 181, sh8 163, sh9 171, sh10 108. |
|
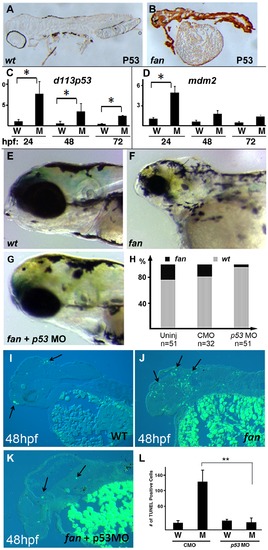
Inhibition of p53 signaling can partially rescue fan mutant craniofacial defects. (A–B) Whole mount IHC analysis of p53 expression in wild type (A) and fan mutants (B). (C, D) qRT-PCR analysis of d113p53 (C) and mdm2 (D) gene in wild type and fan mutants at 24, 48 and 72 hpf. (E–G) Bright field images of live 3 dpf wild type (E), fan mutant (F), and fan mutant injected with p53MO (G). (H) Percent zebrafish exhibiting normal eye development in uninjected (Uninj), control MO (CMO), and p53MO injected embryos. (I–K) TUNEL stained and sectioned wild type (WT) (I), fan mutant (J), and fan mutant injected with p53 MO (K). Arrows indicate apoptotic cells. (L) Statistical analyses revealed significantly reduced apoptosis in fan mutants injected with p53MO. (P value<0.01). |
|
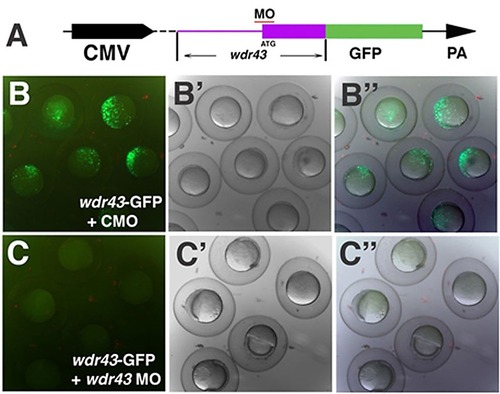
Knockdown efficiency of wdr43 Morpholino. (A) Diagram of the wdr43 reporter construct used to test wdr43 MO knockdown efficiency. GFP (green box) was fused in frame to the 32 end of a portion of the 52 end of the wdr43 cDNA including the 52 UTR (purple line) and first two exons of wdr43 gene (purple box). Red line indicates the MO target region. The chimeric gene was driven by the CMV promoter and followed by the 32 SV40 polyA signal (PA). (B–B3) Fluorescence of shield stage zebrafish embryos injected with the wdr43-GFP reporter construct and control MO (CMO) (B, fluorescent microscopy; B2 bright field, B3 merged fluorescent and bright field). (C–C) Lack of fluorescence in shield stage zebrafish embryos injected with wdr43-GFP reporter constructs and wdr43 MO (C, fluorescent microscopy; C2 bright field; C3 merged fluorescent and bright field). |
|
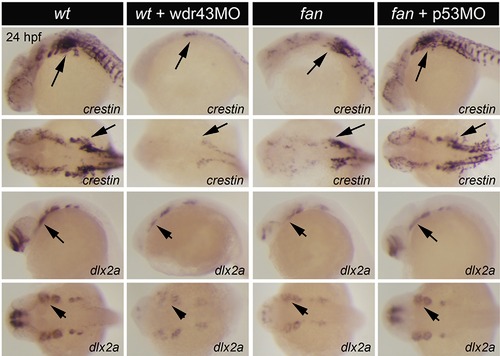
WISH analyses of wdr43 MO injected embryos, and p53 MO injected fan mutants. WISH was performed to examine NCC markers crestin and dlx2a expression in 24 hpf wild type embryos (WT), wild type embryos injected with wdr43 MO, fan mutants, and fan mutants injected with 53 MO, as indicated. wdr43 MO injected embryos exhibited down regulated expression of all NCC markers, similar to that observed in fan mutants (arrows). Injection of p53MO into single cell stage fan mutants rescued NCC marker gene expression (arrows). |
|
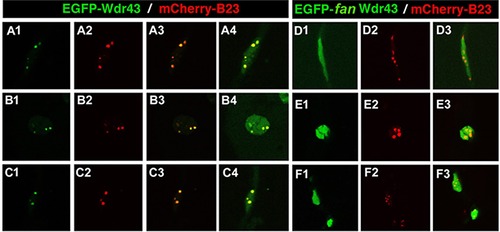
Subcellular localization of EGFP-tagged wild type or fan mutant Wdr43 in zebrafish embryos. Confocal images taken from 24 hpf old zebrafish embryos injected at single cell stage with EGFP tagged wild type (A1–4, B1–4, C1–4) or fan mutant (D1–3, E1–3, F1–3)) wdr43 mRNA. Co-injection of mCherry tagged zebrafish B23 mRNA was used to label nucleoli (A2, B2, C2, D2, E2, F2). Each row of panels indicates different types of cells imaged in the whole embryo. GFP expression was manually saturated in panels (A4, B4, C4) to reveal the entire nucleus. |
|
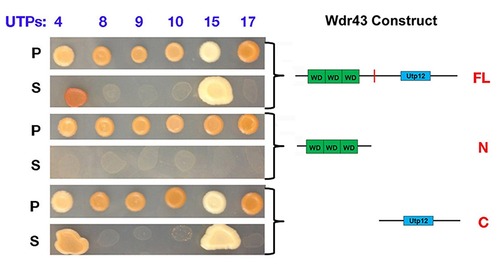
Yeast two hybrid analysis of yeast Wdr43/UTP5 and yeast t-UTP subcomplex proteins. Primers for full length, and fan mutant truncated yeast UTP5 were used to amplify and sublcone these yeast cDNA constructs into pGADT7 vector. The remaining yeast expression constructs were obtained from Dr. S. Baerga. P: permissive medium (-Leu and - Trp). S: selective medium (-Ade, -His, -Leu and -Trp). |