- Title
-
DNA methyltransferase 1 functions through C/ebpa to maintain hematopoietic stem and progenitor cells in zebrafish
- Authors
- Liu, X., Jia, X., Yuan, H., Ma, K., Chen, Y., Jin, Y., Deng, M., Pan, W., Chen, S., Chen, Z., de The, H., Zon, L.I., Zhou, Y., Zhou, J., Zhu, J.
- Source
- Full text @ J. Hematol. Oncol.
|
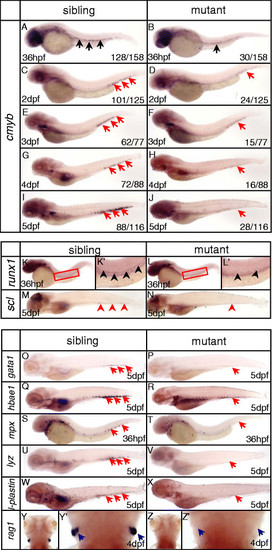
Impairment of definitive hematopoiesis in dnmt1 mutant zebrafish. (A-J) WISH analysis of cmyb expression from 36 hpf to 5 dpf. In ldd794 mutant, cmyb expression was decreased from 36 hpf (A-B) and absent at 5 dpf (I-J). (K, K′, L, L′) Expression of hematopoietic progenitor marker runx1 was decreased at 36 hpf. (K′, L′) Magnified images of the boxed regions in K and L, respectively. (M-N) Expression of hematopoietic progenitor marker scl was decreased at 5 dpf. (O-Z) WISH analysis of key hematopoietic markers in dnmt1 mutant embryos and wild-type siblings. Expression of erythrocyte progenitor marker gata1 (O-P), mature erythrocyte marker hbae1(Q, R), myeloid-specific marker mpx (S, T), macrophage marker lyz (U, V), l-plastin (W, X) were decreased. Expression of lymphocyte maker rag1 (Y, Y′, Z, Z′) was completely absent at 4 dpf. Blue arrows indicate the position of the thymus; red arrows indicate the CHT; black arrows indicate the AGM. |
|
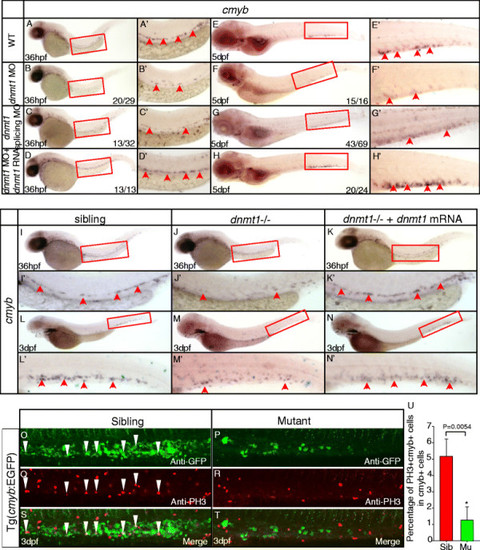
Dnmt1 mRNA rescue and anti-phosphorylated histone H3 (pH3) immunostaining assays in dnmt1-deficient embryos. (A-H) Dnmt1 mRNA rescue assay at 36 hpf (A-D) and 5 dpf (E-H). Dnmt1 ATG morpholino (4 ng) and splicing-specific morpholino (8 ng) effectively reproduced decreased cmyb expression phenotype in wild-type embryos (B, C, F, G). Dnmt1 mRNA rescued the defects of dnmt1 morphants. (I-N) Rescue assay of wild-type dnmt1 mRNA in ldd794 dnmt1 mutants at 36 hpf (I-K) and 3 dpf (L-N). Wild-type dnmt1 mRNA effectively restored cmyb expression in ldd794 dnmt1 mutants at 36 hpf (K) and 3 dpf (N). (A′-N′) Magnified images of the boxed regions in A to N, respectively. Red arrows indicate cmyb-positive HSPCs in the AGM or CHT. (O-T) The proliferation of HSPCs was reduced in dnmt1/ mutants. Double immunostaining of cmyb-EGFP (O-P) and anti-PH3 (Q-R) in the CHT of Tg(cmyb-EGFP) line at 72 hpf. The bottom panel shows merged images (S, T). (U) Statistics of PH3 and cmyb-EGFP double positive cells. |
|
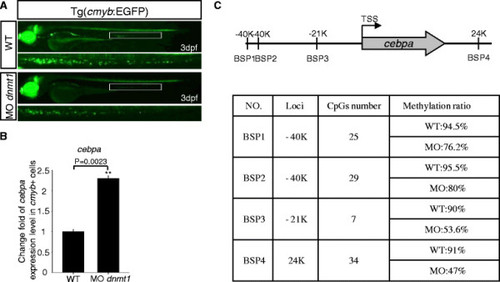
Fluorescent images of the Tg(cmyb-EGFP) line, Q-PCR result of cebpa expression, and bisulfite sequencing PCR (BSP) assay. (A) Fluorescent images of the Tg (cmyb-EGFP) line at 3 dpf. EGFP positive cells in Tg (cmyb:EGFP) dnmt1 morphants were sharply decreased compared with wild-type ones. (B) Cebpa expression level in wild-type and dnmt1 knockdown cmyb-EGFP positive cells. Q-PCR result indicated that there was a significant increase of cebpa expression level in dnmt1 morphants compared with wild-type ones. (C) BSP analysis of putative cebpa regulation regions in cmyb-EGFP positive cells at 3 dpf. The methylation levels of cebpa regulation regions in dnmt1 morphants were significantly decreased in all the four chosen regions compared to those in wild-type embryos. EXPRESSION / LABELING:
PHENOTYPE:
|
|
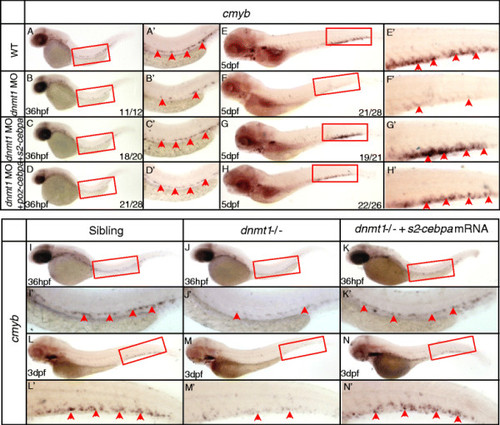
SUMO2-C/ebpa, POZ-C/ebpa rescue assay in dnmt1 morphants and ldd794 dnmt1 mutants. (A-H) SUMO2-C/ebpa and POZ-C/ebpa restored cmyb expression in dnmt1 MO knockdown morphants at 36 hpf (A-D) and 5 dpf (E-H). (I-N)SUMO2-C/ebpa mRNA restored cmyb expression in the AGM region at 36 hpf (K) and the CHT region at 3 dpf (N) of ldd794 dnmt1 mutants. (A′-N′) Magnified images of the boxed regions in A to N. Red arrows indicate cmyb-positive HSPCs in the AGM or CHT. |
|
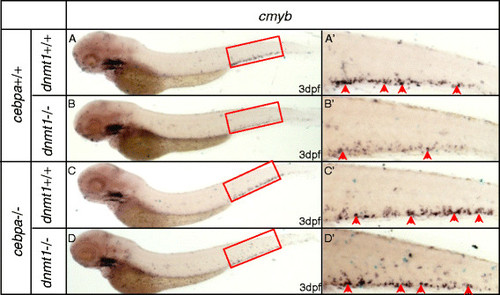
WISH assays of cmyb in cebpa null mutants and siblings with or without dnmt1 mutation. (A) WISH assay of cmyb in embryos without cebpa and dnmt1 homozygote mutation. (B) WISH assay of cmyb in embryos only with dnmt1 homozygote mutation. Note that cmyb was markedly decreased. (C) WISH assay of cmyb in embryos only with cebpa homozygote mutation. (D) WISH assay of cmyb in embryos both with cebpa and dnmt1 homozygote mutation. Note that cmyb was comparable with embryos without dnmt1 homozygote mutation. (A′-D′) Magnified images of the boxed regions in A to D. Red arrows indicate cmyb-positive HSPCs in the CHT. EXPRESSION / LABELING:
PHENOTYPE:
|
|
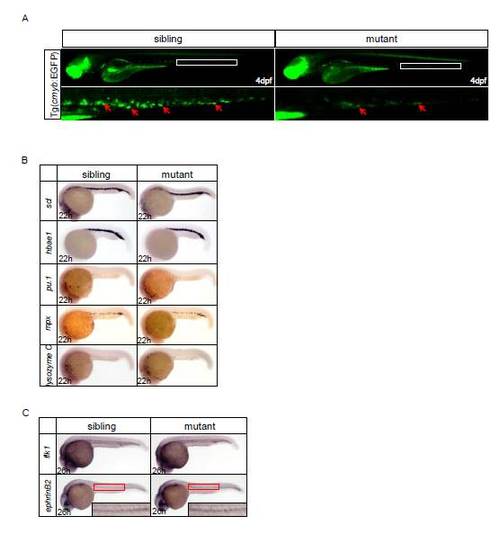
WISH assays of hematopoietic markers of ldd794. (A) EGFP positive cells in Tg(cmyb:EGFP) dnmt1 mutant embryo were sharply decreased compared with wild-type siblings. (B) The primitive hematopoietic markers of mutant embryos were comparable with siblings at 22hpf. (C) The vascular markers flk1 and ephrinB2 were normal in ldd794 mutant. |
|
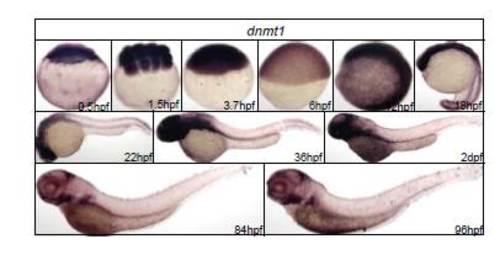
WISH analysis of dnmt1. Zebrafish dmnt1 was expressed ubiquitously. |
|
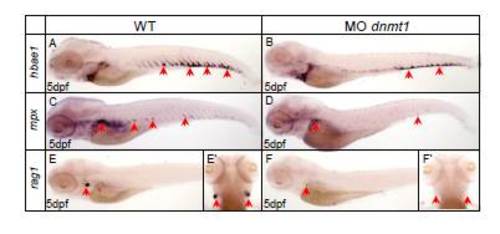
WISH analysis of erythroid, myeloid and lymphoid markers in dnmt1 morphants. The mature erythrocyte marker hbae1 (A, B), myeloid-specific marker mpx (C, D) and lymphoid-specific marker rag1 (E, E', F, F') were all decreased. |
|
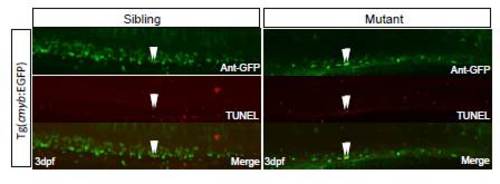
TUNEL assays in sibling and dnmt1-/- mutant. Double positive cells in sibling and dnmt1 mutant were indicated by arrows. |
|
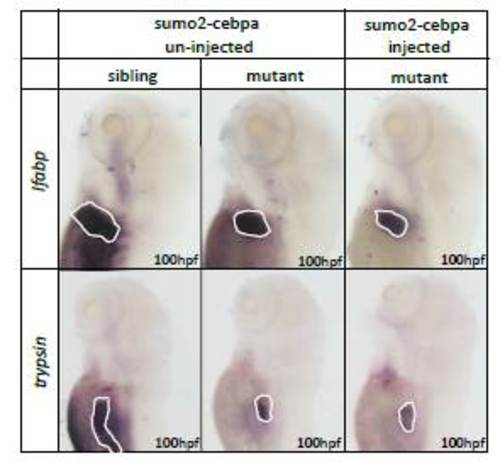
WISH assays of hepatocyte marker lfabp and pancreas marker trypsin. The expressions oflfabp and trypsin were clearly decreased in ldd794 mutants. Note that SUMO2-C/ebpa was unable to rescue the development defects of liver and pancreas. |
|
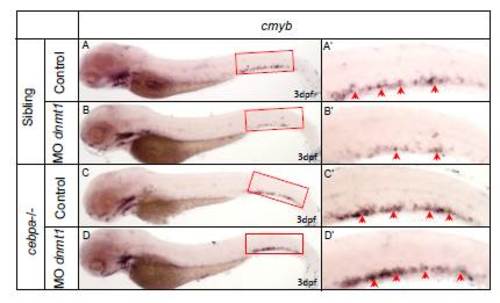
WISH assays of cmyb in cebpa null mutants and siblings with or without dnmt1 knock down. (A) WISH assay of cmyb in siblings. (B) WISH assay of cmyb in siblings injected with dnmt1 morpholino. Note that cmyb was markedly decreased. (C) WISH assay of cmyb in cebpa null mutants. (D) WISH assay of cmyb in cebpa null mutants injected with dnmt1 morpholino. Note that cmyb was comparable with un-injected siblings. (A'-D') Magnified images of the boxed regions in A to D respectively. Red arrows indicate cmyb-positive HSPCs in the CHT. |