- Title
-
Expression and retinoic acid regulation of the zebrafish nr2f orphan nuclear receptor genes
- Authors
- Love, C.E., and Prince, V.E.
- Source
- Full text @ Dev. Dyn.
|
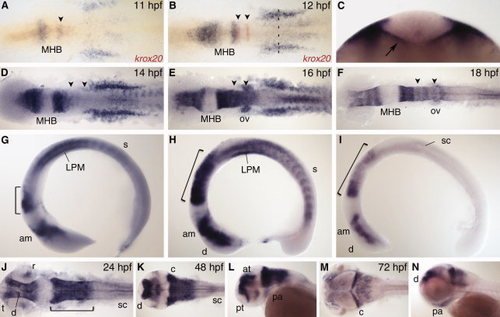
nr2f1a expression. A,B: Dorsal view during early segmentation stages co-stained with r3 and r5 marker krox20 (red). C: Optical section of 12 hours post fertilization (hpf) embryo at anteroposterior level indicated by dashed line in B. Arrow indicates endoderm expression. D–I: Dorsal view (D–F) and lateral view (G–I) during segmentation stages. J,K,M: Dorsal view. L,N: Lateral view. am, anterior midbrain; at, anterior tegmentum; c, cerebellum; d, diencephalon; LPM, lateral plate mesoderm; MHB, midbrain–hindbrain boundary; ov, otic vesicle; pa, pharyngeal arch; pt, prethalamus; r, retina; s, somite; sc, spinal cord; t, telencephalon. Arrowheads indicate r3 and r5. Bracket indicates hindbrain. EXPRESSION / LABELING:
|
|
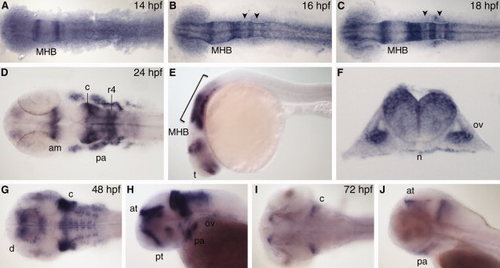
nr2f1b expression. A–C: Dorsal view during segmentation stages, anterior to left. D,G,I: Dorsal view. E,H,J: Lateral view, anterior to left. F: Transverse section through r5. am, anterior midbrain; at, anterior tegmentum; c, cerebellum; d, diencephalon; MHB, midbrain–hindbrain boundary; n, notochord; ov, otic vesicle; pa, pharyngeal arch; pt, prethalamus; r4, rhombomere 4. Arrowheads indicate r3 and r5. Bracket indicates hindbrain. EXPRESSION / LABELING:
|
|
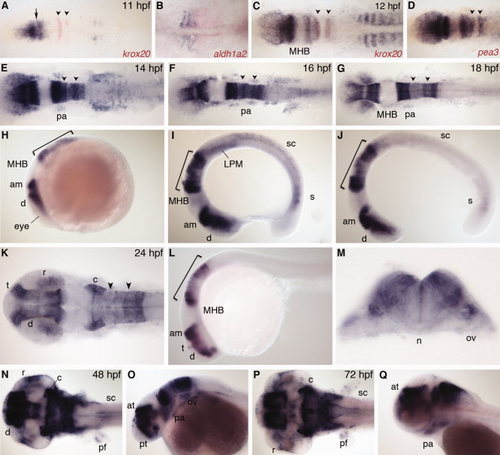
nr2f2 expression. A,C: Dorsal view, co-stained with krox20 (red). Arrow indicates gap between presumptive diencephalon and anterior midbrain. B: Dorsal view of trunk, anterior to left, co-stained with aldh1a2 (red). D: Dorsal view, co-stained with pea3 (red). E–G,K,N,P: Dorsal view. H–J,L,O,Q: Lateral view. M: Transverse section through r5. am, anterior midbrain; at, anterior tegmentum; c, cerebellum; d, diencephalon; LPM, lateral plate mesoderm; MHB, midbrain–hindbrain boundary; n, notochord; ov, otic vesicle; pa, pharyngeal arch; pf, pectoral fin; pt, prethalamus; r, retina; s, somite; sc, spinal cord; t, telencephalon. Arrowheads indicate r3 and r5. Bracket indicates hindbrain. EXPRESSION / LABELING:
|
|
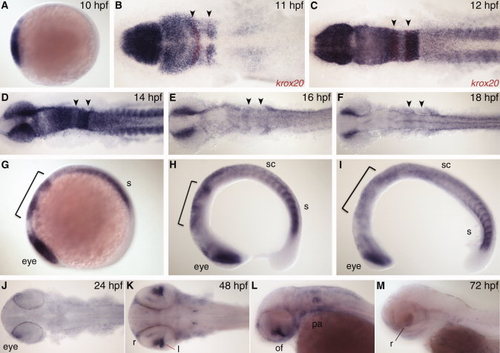
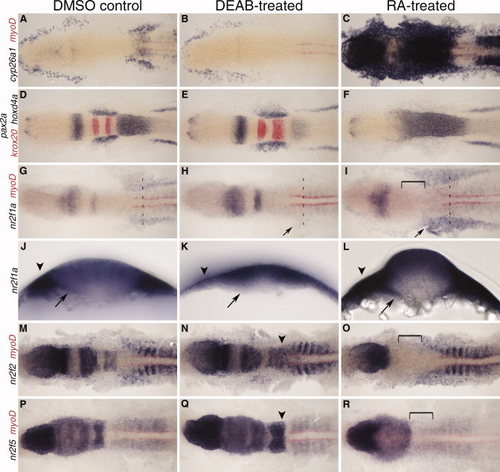
nr2f expression at 12 hours post fertilization (hpf) with retinoic acid (RA) and DEAB treatment 8–10 hpf. A–C: Expression of RA target gene cyp26a1 to confirm treatment efficiency, co-stained with myoD (red) to mark the myotome. D–F: MHB and hindbrain markers pax2a, krox20 (red), and hoxd4a show changes in tissue specification. G–I: nr2f1a expression. Arrow indicates changes in lateral plate mesoderm (LPM) expression. J–L: Transverse sections showing changes in nr2f1a transcript levels in the endoderm (arrows) and LPM (arrowheads) at anteroposterior levels designated by dashed lines in G–I. M–O: nr2f2 expression. P–R: nr2f5 expression. N,Q: Arrowheads indicate a posterior extension of hindbrain expression. Brackets indicate loss of hindbrain expression. EXPRESSION / LABELING:
|
|
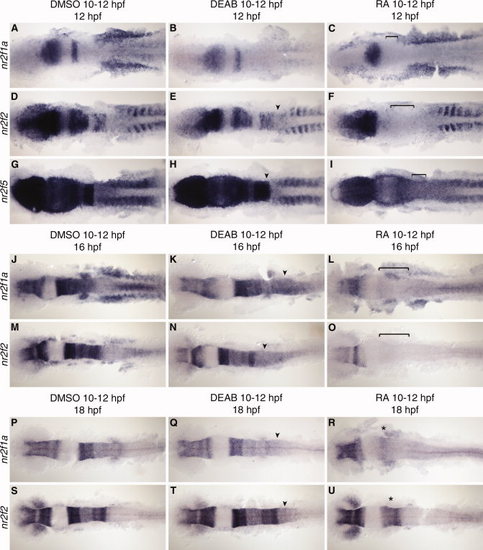
nr2f expression at 12, 16, and 18 hours post fertilization (hpf) with retinoic acid (RA) and DEAB treatment 10–12 hpf. A–I: nr2f1a, nr2f2, and nr2f5 expression at 12 hpf. J–O: nr2f1a and nr2f2 expression at 16 hpf after RA or DEAB treatment from 10–12 hpf. P–U: nr2f1a and nr2f2 expression at 18 hpf after RA or DEAB treatment from 10–12 hpf showing a return of anterior hindbrain expression (asterisk). Arrowheads indicate a posterior extension of hindbrain expression. Brackets indicate loss of hindbrain expression. EXPRESSION / LABELING:
|
|
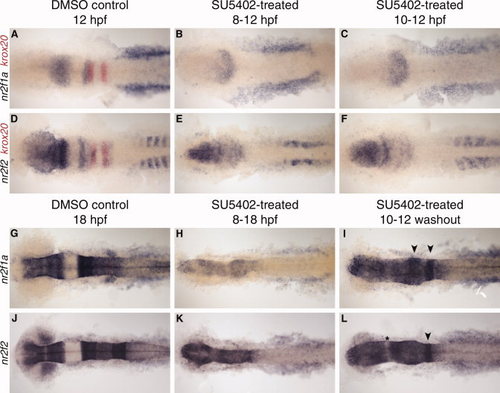
nr2f expression at 12 and 18 hours post fertilization (hpf) after SU5402 treatment. A–F: nr2f1a and nr2f2 expression at 12 hpf. SU5402 treatment from 8–12 hpf (B,E) or 10–12 hpf (C,F). G–L: nr2f1a and nr2f2 expression at 18 hpf. SU5402 treatment from 8–18 hpf (H,K) or 10–12 hpf and washed out (I,L). Arrowheads indicate rhombomere-specific expression in the hindbrain. Asterisk indicates gap in expression, possibly between midbrain and hindbrain. EXPRESSION / LABELING:
|