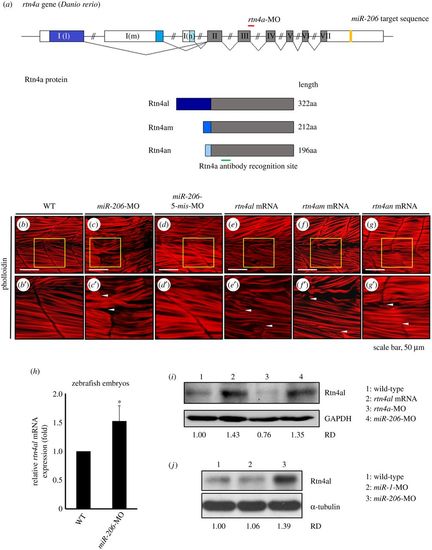
Knockdown of endogenous miR-206 increases the level of Rtn4al protein, causing abnormal transverse actin filaments across the somite boundary. (a) The genomic structure of three isoforms of rtn4a genes. Locations of the rtn4a-MO binding sequence (red line), miR-206 binding site at the 3′UTR of rtn4a mRNA (yellow box), and recognized region of antibody against Rtn4a protein (green line) were indicated. (b) WT, (c) miR-206-MO-injected, (d) miR-206-5-mis-MO-injected, (e) rtn4al-mRNA-injected, (f) rtn4am-mRNA-injected, and (g) rtn4an-mRNA-injected embryos developed at 48 hpf were immunostained with fluorescent phalloidin to label F-actin. Panels (b′–g′) were the enlarged views of corresponding panels (b–g). The area between the two arrowheads indicates the abnormal transverse actin filaments across the somite boundary. (h) The relative amounts of rtn4al mRNA between WT deheaded embryos and miR-206-MO-injected deheaded embryos at 20 hpf were quantified by qPCR when that of WT was normalized as 1. One hundred embryos were used each time, and experiments were performed in triplicate (n = 3). (i) Western blot analysis of Rtn4al (35 kD) among proteins extracted from deheaded embryos at 20 hpf: in WT (lane 1), rtn4al-mRNA-injected embryos (lane 2), rtn4a-MO-injected embryos (lane 3) and miR-206-MO-injected embryos (lane 4). (j) Western blot analysis of Rtn4al among proteins extracted from deheaded embryos at 20 hpf: in WT (lane 1), miR-1-MO-injected embryos (lane 2), and miR-206-MO-injected embryos (lane 3). Student's t-test was used for statistical analysis. Asterisk indicates significant difference at *p < 0.05. RD: Relative density. GAPDH and tubulin served as internal controls.
|

