- Title
-
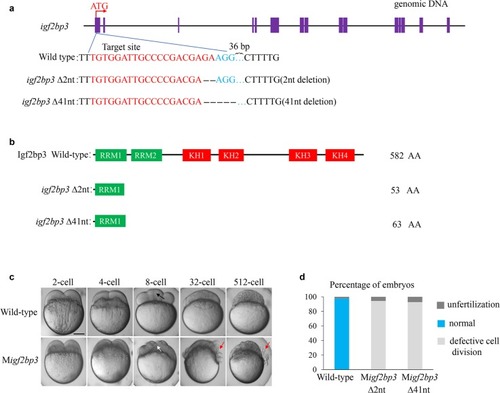
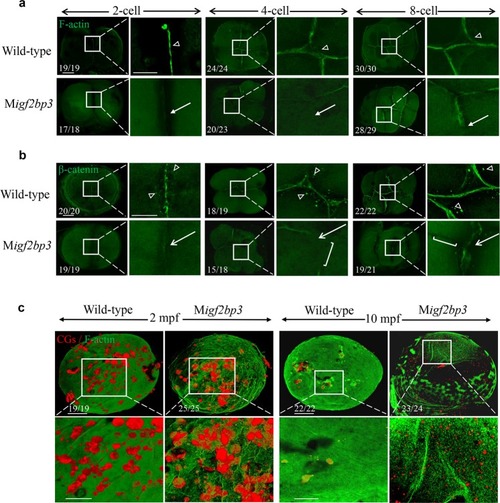
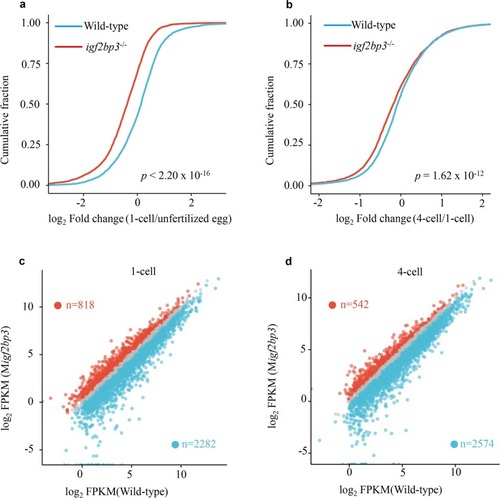
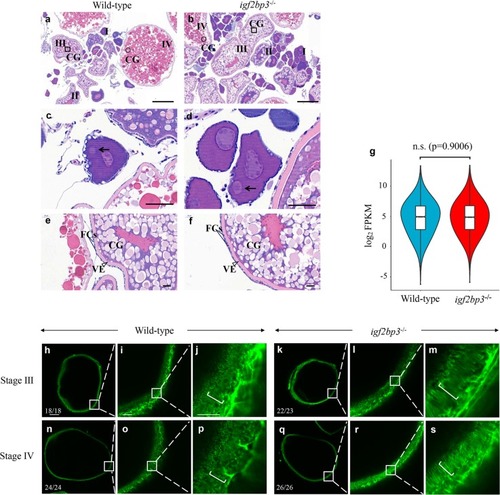
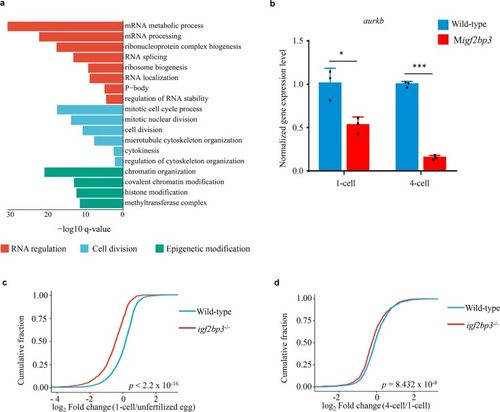
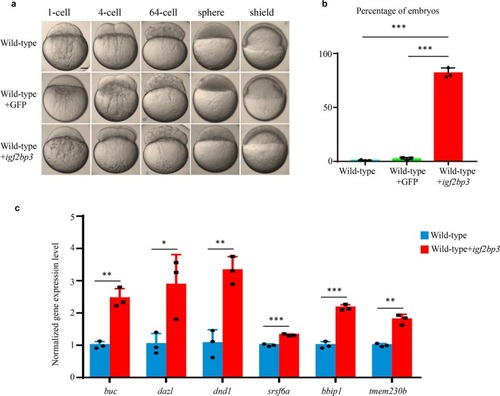
Igf2bp3 maintains maternal RNA stability and ensures early embryo development in zebrafish
- Authors
- Ren, F., Lin, Q., Gong, G., Du, X., Dan, H., Qin, W., Miao, R., Xiong, Y., Xiao, R., Li, X., Gui, J.F., Mei, J.
- Source
- Full text @ Commun Biol
|
|
|
|
|
|
|
PHENOTYPE:
|
|
|
|
|

Unillustrated author statements PHENOTYPE:
|