- Title
-
Using Zebrafish to Study Collective Cell Migration in Development and Disease
- Authors
- Olson, H.M., Nechiporuk, A.V.
- Source
- Full text @ Front Cell Dev Biol
|
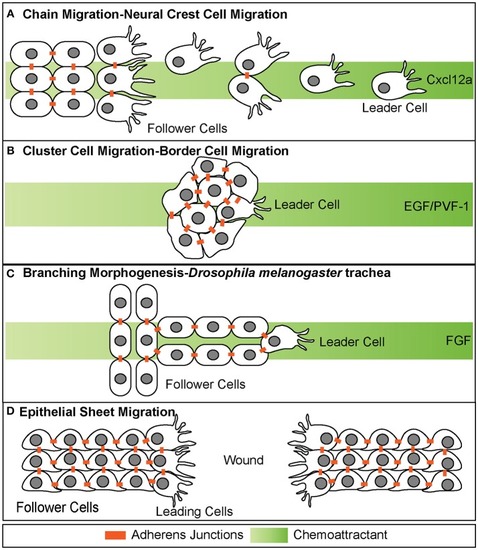
Different modes of collective cell migration. |
|
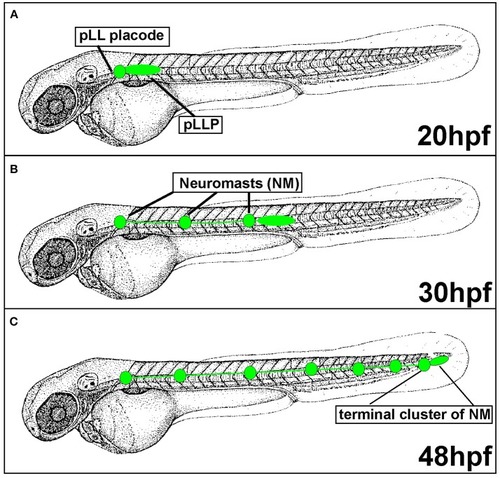
Posterior Lateral Line formation (pLL) and posterior Lateral Line Primordium (pLLP) migration. |
|
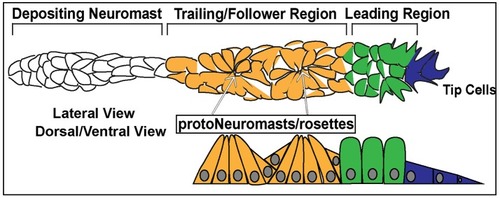
Schematic of the posterior Lateral Line Primordium (pLLP). In blue are the 2–3 leader cells. In green is the leading region. In orange is the trailing region. In white is a depositing neuromast. The top schematic is a lateral view of pLLP cells. The bottom schematic is a dorsal/ventral view. Arrows point to proto-neuromasts/rosettes. |
|
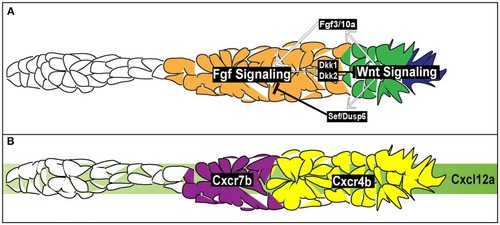
Signaling and chemotactic pathways active during pLLP migration. |
|
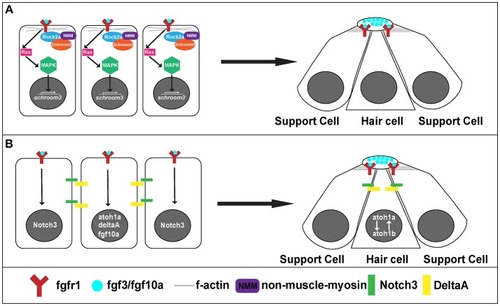
Signaling pathways for rosette formation and proto-NM maturation. |
|
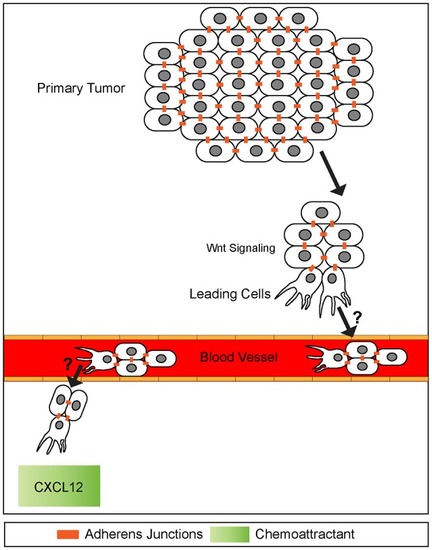
A potential model of collective cell invasion. Small groups of cells high in canonical Wnt signaling break off from the primary tumor, invade surrounding tissue, and enter the blood stream. This cluster then extravisates from the blood stream homing onto to a chemokine source, such as CXCL12. The mechanisms of blood vessel intravisation and extravasation by cell clusters are not known (question marks). It has been speculated that behaviors of such clusters exhibit many similarities to leading front cells present in collective cell migration during embryogenesis. Adherens junctions are maintained throughout invasion process. |