Fig. 2
- ID
- ZDB-FIG-060119-5
- Publication
- Alioto et al., 2005 - The odorant receptor repertoire of teleost fish
- Other Figures
- All Figure Page
- Back to All Figure Page
|
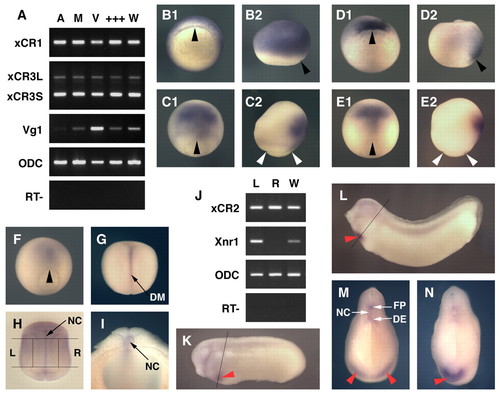
Spatial expression patterns of Xenopus EGF-CFC genes. (A) Spatial expression patterns of XCR1, XCR3S and XCR3L at stage 9, as determined by RT-PCR of the animal region (A), the marginal region (M), the vegetal region (V), a mixture of the dissected three parts (+++) and intact whole embryos (W). VG1 was used as a control for dissection. (B-I,K-N) Spatial expression analyses by in situ hybridization for XCR1 (FRL1; B,C), XCR3 (D,E) and XCR2 (F-I,K-N). (B1,D1) Stage 10+ in vegetal view. The dorsal side is up. (C1,E1,F,G) Stage 12 (C,E,F) and stage 15 (G) in dorsal view. The anterior side is up. (B2,C2,D2,E2) Stage 10+ (B,D) and stage 12 (C,E) in lateral view. The anterior side is up and the dorsal side is to the right. (H) Stage 20 in dorsal view. Embryo was cleared with benzyl benzoate/benzyl alcohol. Lines show the left (L) and right (R) dissection model for RT-PCR (J). (I) Transverse bisected embryo at stage 20. (J) RT-PCR of the left lateral region (L), the right lateral region (R) and intact whole embryos (W) at stage 20. XCR2 was expressed bilaterally, whereas XNR1 was expressed on only the left side. (K) Stage 25. Line indicates the section plane shown in M. (L) Stage 30. Line indicates the section plane in N. (M,N) Transverse cut embryos at stage 25 (M) and stage 30 (N). Red arrowheads indicate the expression of XCR2 in the prospective heart region; black arrowheads indicate the dorsal lip; white arrowheads indicate the blastopore. DE, dorsal endoderm; DM, dorsal midline; FP, floorplate; NC, notochord. |