- Title
-
Genetic Mechanism That Defines the Characteristic Neurogenesis Patterns in the Neural Plate Using Hes/her Genes During Early Vertebrate Development
- Authors
- Ohyanagi, T., Tokizaki, H., Sato, T., Tsuruoka, M., Yamasu, K.
- Source
- Full text @ Genesis
|
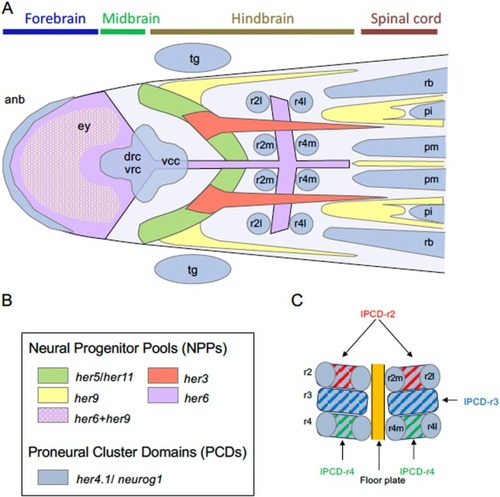
Summary of the neurogenesis pattens in the developing neural plate in zebrafish. (A) Schematic diagram of the anterior neural plate, showing distribution of proneural cluster domains (PCDs) and neural progenitor pools (NPPs) marked by various |
|
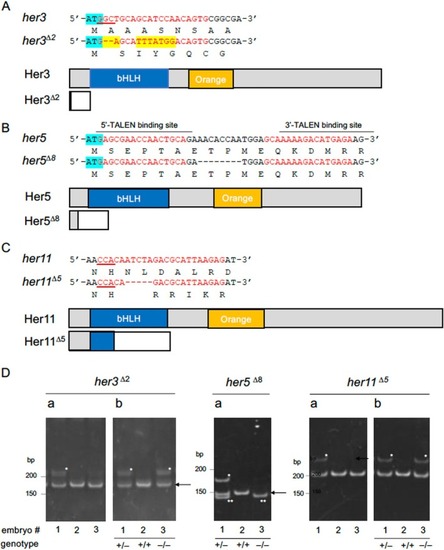
Structures of the mutant |
|
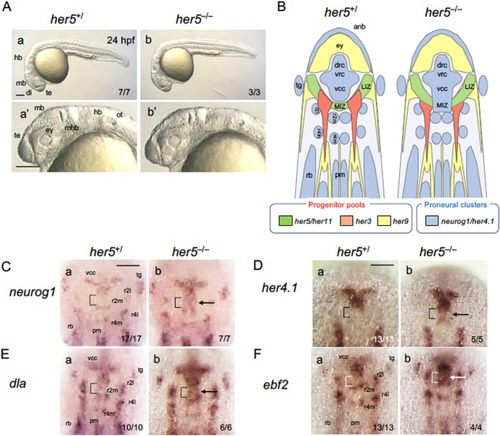
Phenotypes of |
|
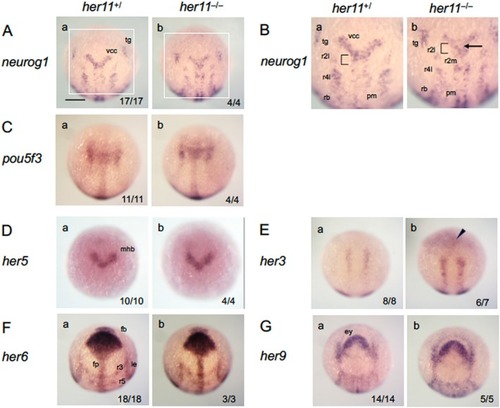
Expression of brain regionalization genes and |
|
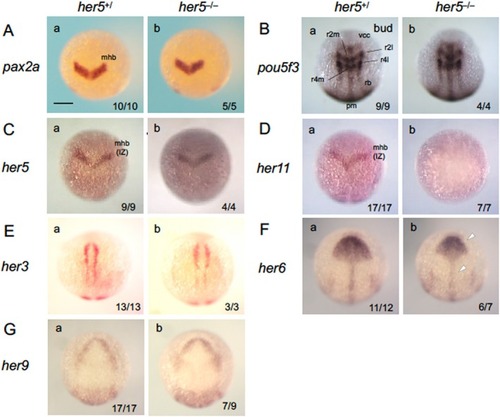
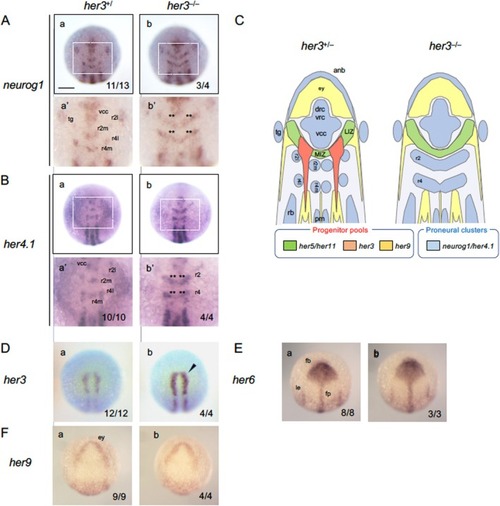
Phenotypic analysis of |
|
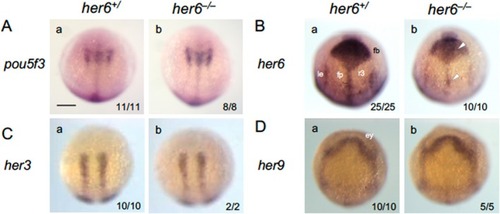
Phenotypic analysis of |
|
Expression of Notch‐independent |
|
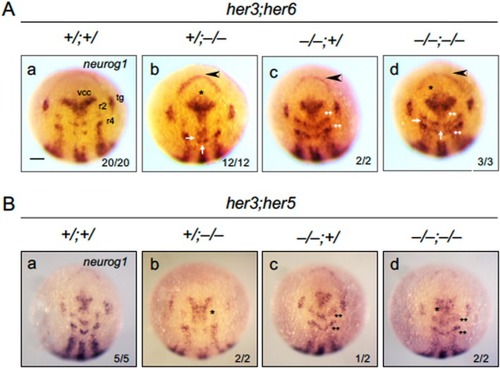
Abnormal neurogenesis in |