- Title
-
Efficient targeted integration directed by short homology in zebrafish and mammalian cells
- Authors
- Wierson, W.A., Welker, J.M., Almeida, M.P., Mann, C.M., Webster, D.A., Torrie, M.E., Weiss, T.J., Kambakam, S., Vollbrecht, M.K., Lan, M., McKeighan, K.C., Levey, J., Ming, Z., Wehmeier, A., Mikelson, C.S., Haltom, J.A., Kwan, K.M., Chien, C.B., Balciunas, D., Ekker, S.C., Clark, K.J., Webber, B.R., Moriarity, B.S., Solin, S.L., Carlson, D.F., Dobbs, D.L., McGrail, M., Essner, J.
- Source
- Full text @ Elife
|
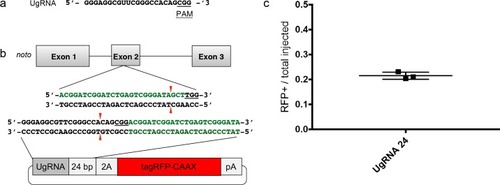
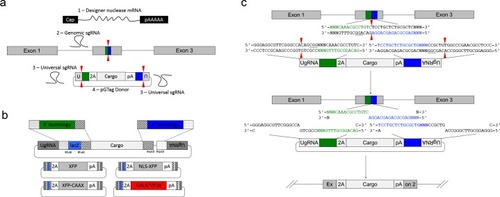
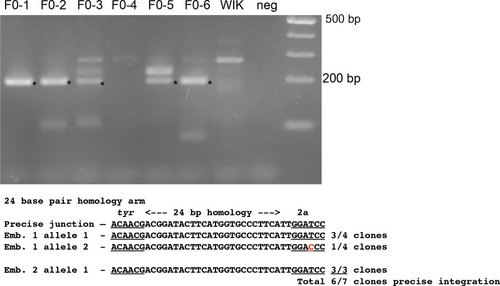
A single short homology arm 5’ to the sgRNA target site in the noto gene targets integration in zebrafish embryos. (a) Schematic for noto homology arm and donor vector design. Bold letters show the noto sgRNA target sequence in the genome. This sgRNA target sequence was also used to target Cas9 cutting in the donor vector. Black bars represent the different homology arm lengths 12, 24, or 48 bp, used to target the 2A-tagRFP-CAAX donor vector into the noto exon 1 target site. PAM sequences are underlined. Red arrows indicate the Cas9 cut site 3 bp upstream of the PAM. The 3 nucleotide spacer lacking homology to the genome is represented by the lowercase sequence ‘aaa’ located in between the donor vector PAM and the 5’ end of the homology arm. (b) Targeting efficiency of noto exon1 2A-tagRFP-CAAX donor vectors containing a single 5’ homology arm of 12, 24, or 48 bp. Data represents mean +/- s.e.m. of 3 independent targeting experiments. p values calculated using two-tailed unpaired t-test. (c) Live confocal image of noto-2A-TagRFP-CAAX-SV40 targeted embryo showing specific RFP expression in the notochord. Scale bar, 100 μm. (d) Sanger sequencing of cloned 5’ junction fragments from RFP positive F0 embryos, aligned to the expected sequence from a precise integration event. Numerator represents correct clones, denominator represents total clones sequenced. Junctions are considered precise if the homology arm does not contain any mismatch and there are no insertions or deletions up- or downstream of the programmed homology. |
|
( |
|
( |
|
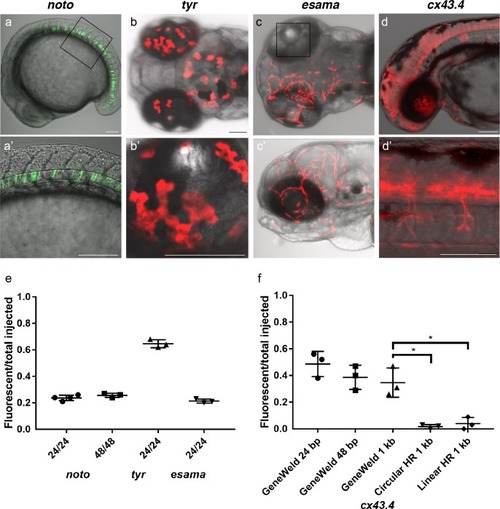
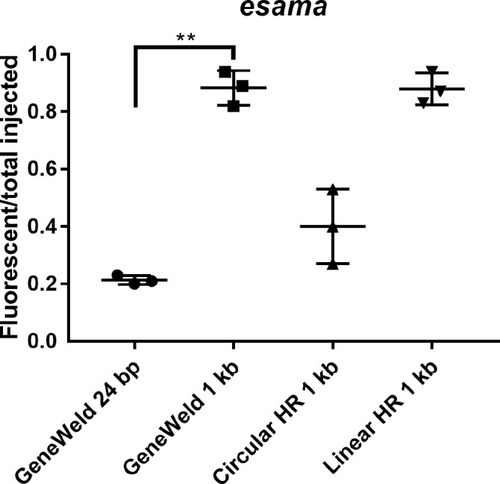
(a–d) Live confocal images of F0 injected embryos showing fluorescent reporter expression after GeneWeld targeted integration. (a, a’) Mid somite stage embryo targeted at noto with 2A-eGFP. (b, b’) 5 days post fertilization (dpf) Tg(UAS:mRFP)tpl2 embryo targeted at tyr with 2A-Gal4/VP16. (c) 2 dpf and (c’) 3dpf Tg(UAS:mRFP)tpl2 embryo targeted at esama with −2A-Gal4/VP16. (d, d’) 31 hr post fertilization embryo targeted at cx43.4 with 2A-tagRFP-CAAX. (e) Fraction of embryos with reporter gene expression following GeneWeld targeting at noto, tyr and esama. 5’ and 3’ homology lengths flanking donor cargos indicated in base pairs as 24/24 or 48/48. (f) Comparison of the fraction of RFP expressing embryos after targeting cx43.4 exon 2 using GeneWeld 24/24 bp homology, GeneWeld 48/48 bp homology, Geneweld 1 kb/1 kb homology, Circular HR 1 kb/1 kb (injection did not include UgRNA, *p=0.0067), Linear HR 1 kb/1 kb (donor was digested and the linear DNA fragment containing the homology arm targeting construct was gel purified before injection, *p=0.0111). Data represents mean +/- s.e.m. of 3 independent targeting experiments. p values calculated using Students t test. Scale bars, 100 μm. |
|
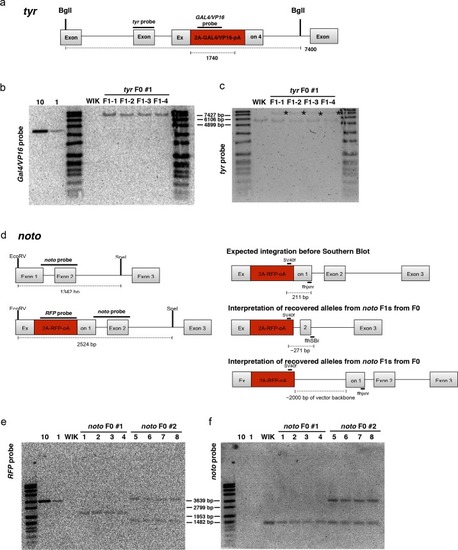
PCR amplification and sequence of 5’ junction fragments between |
|
Comparison of the frequency of RFP expressing injected embryos after targeting |
|
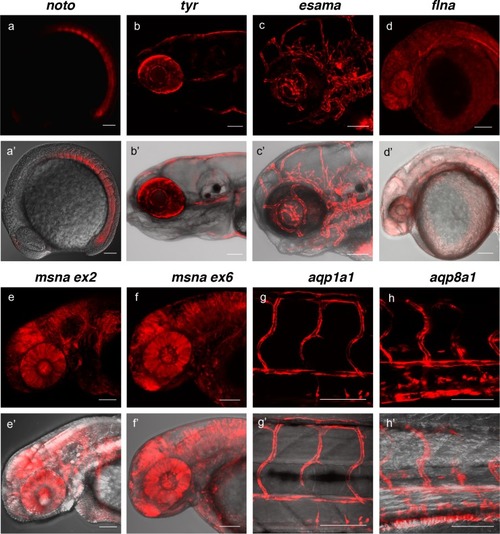
(a, a’) Tg(noto-2A-TagRFP) embryo at mid somite stage showing expression in the notochord and floor plate. (b, b’) Tg(tyr-2A-Gal4/VP16); Tg(UAS:mRFP)tpl25 dpf larva displaying expression in the melanocytes. (c, c’) Tg(esama-2A-Gal4/VP16); Tg(UAS:mRFP)tpl24 dpf larva showing expression in the vascular system. (d, d’) Tg(flna-2A-Gal4/VP16); Tg(UAS:mRFP)tpl21 dpf embryo showing widespread expression. (e, e’ and f, f’) Exon 2 and exon 6 msna targeted Tg(msna-2A-Gal4/VP16); Tg(UAS:mRFP)tpl2 2dpf embryos showed expression in the central nervous system and vasculature. (g, g’ and h, h’) Tg(aqp1a1-2A-Gal4/VP16; Tg(UAS:mRFP)tpl2) and Tg(aqp8a1-2A-Gal4/VP16); Tg(UAS:mRFP)tpl22 dpf embryos display RFP expression in the trunk and tail vasculature. Scale bars, 100 μm. |
|
( |
|
Precise integration at the 5’ and 3’ ends in F1 progeny from F0 founder fish targeted at |
|
|
|
|
|
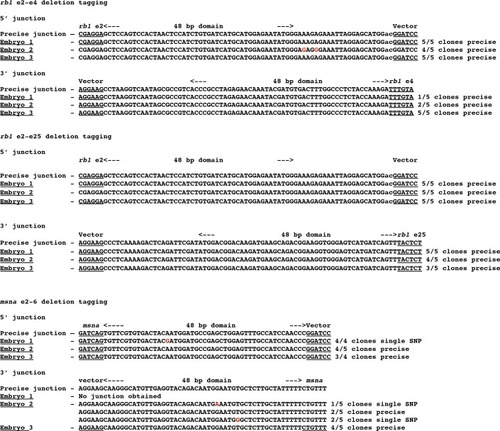
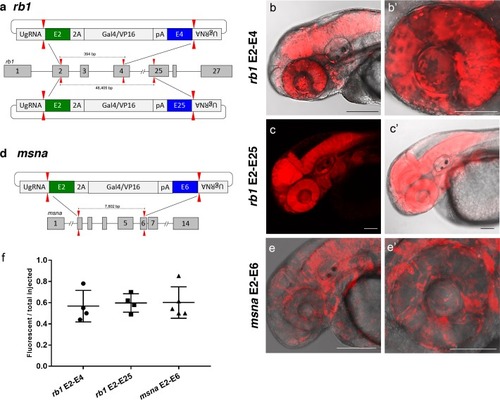
(a) Schematic for Gal4/VP16 reporter integration to tag a deletion allele of rb1 exons 2–4 (top) and rb1 exons 2–25 (bottom). Arrowheads designate CRISPR/Cas9 DSBs. CRISPR sgRNAs in two exons are expected to excise the intervening genomic DNA. The targeting vector contains a 5’ homology arm flanking the upstream exon target site and a 3’ homology arm flanking the downstream exon target site. (b, b’) Live confocal image of F0 Tg(UAS:mRFP)tpl2 embryo after 2A-Gal4/VP16 deletion tagging at rb1 exons 2–4. (c, c’) Live confocal image of F1 Tg(rb1-e2-2A-Gal4/VP16) embryo from a founder targeted at rb1 exons 2–25. A deletion from exon 2–25 was not observed in the F1 generation, but the 5’ junction was in frame. (d) Schematic for 2A-Gal4/VP16 deletion tagging of msna exons 2–6. (e, e’) Live confocal image of F0 Tg(UAS:mRFP)tpl2 embryo after 2A-Gal4/VP16 deletion tagging at msna exons 2–6. (f) Somatic reporter efficiency of targeted deletion tagging using 48 bp homology arms for rb1 exons 2–4, rb1 exons 2–25, and msna exons 2–6. Data represents mean +/- s.e.m. of 4 (rb1) and 5 (msna) independent targeting experiments. Scale bars 200 μm (b, c, c’, e); 100 μm (b’, e’). |
|
Detection of precise and imprecise 5’ and 3’ junction fragments in somatic tissue of F0 embryos injected with two guides that target two exons and a pGTag-Gal4/VP16 donor with 5’ and 3’ homology arms corresponding to the 5’ exon and 3’ exon target sites. Cloned PCR amplicons were sequenced from 3 individual embryos for each targeted deletion tagging experiment. |
|
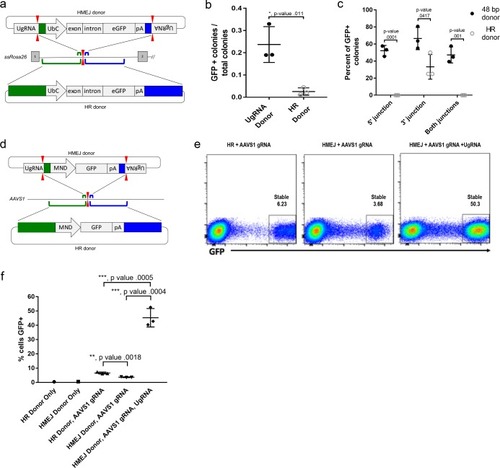
(a) Strategy for integration using HMEJ and HR donors into intron 1 of S. scrofa ROSA26 locus. Arrowheads CRISPR/Cas9 (for HMEJ donor) and TALEN (genome) DSBs. (b) Targeting efficiency of the HMEJ donor vs the HR donor as reported by GFP positive colonies out of total colonies. (c) Percent of GFP positive colonies analyzed containing properly sized junction fragments, comparing HMEJ and HR donors. Data are from three independently targeted cell populations. Data represents mean +/- s.e.m. of 3 independent targeting experiments. (d) Diagram of HR and HMEJ strategies for targeted integration of a MND:GFP reporter cassette into the human AAVS1 locus. (e) Flow cytometry analysis of GFP expression 14 days post-electroporation for each targeting modality: HR (left), HMEJ without universal sgRNA (middle), and HMEJ with universal sgRNA (right). Stable gate was drawn to measure the uniformly expressing population formed by targeted integration and was set based on episome only controls. (f) Quantification of stable GFP expressing population as measured by flow cytometry at day 14. Data are from three independently targeted cell populations. Data represents mean +/- s.e.m. of 3 independent targeting experiments. p values calculated using two-tailed unpaired t-test. |
|
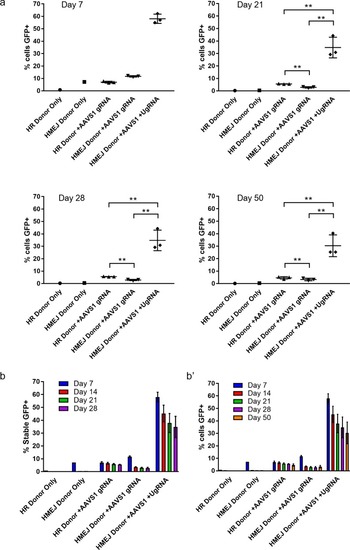
FACs sorted percent of GFP+ cells out of total K-562 cells at day 7, 21, 28, and 50. (b) Summary data for percent of stable GFP+ K-562 cells from day 7, 14, 21, and 28. (b’) Summary data for percent of total cells GFP+ from day 7, 14, 21, 28, and 50. Data represents mean +/- s.e.m. of 3 independent targeting experiments. p values calculated using two-tailed unpaired t-test. |
|
( |