- Title
-
Whole Genome Sequencing-Based Mapping and Candidate Identification of Mutations from Fixed Zebrafish Tissue
- Authors
- Sanchez, N.E., Harty, B.L., O'Reilly-Pol, T., Ackerman, S.D., Herbert, A.L., Holmgren, M., Johnson, S.L., Gray, R.S., Monk, K.R.
- Source
- Full text @ G3 (Bethesda)
|
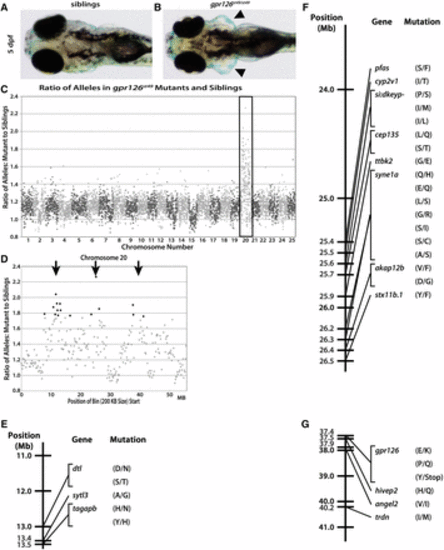
The st49 allele is accurately mapped to chromosome 20 and gpr126 using gDNA extracted from fresh tissue. Dorsal views of phenotypically WT (A) and gpr126st49/st49 (B) zebrafish show the characteristically enlarged ears (arrowheads) of mutants at 5 dpf. (C) When the ratio of variant to reference alleles in the mutant pool is compared to the sibling pool and graphed across the whole genome for gpr126st49, there is a clear spike on chromosome 20 for the gpr126st49 mutation (box). This spike indicates linkage of the region to the trait used to sort the mutant and sibling pools. (D) gpr126st49 is linked to three separate regions on chromosome 20 (arrows). (E–G) Linkage map of the gpr126st49 allele showing the three different regions of chromosome 20 linked to the expanded ear phenotype that was used to sort the mutant and sibling pools. Between all three linked regions there are 29 different protein-coding, homozygous, nonsynonymous SNPs. The single introduced stop is the mutation responsible for the gpr126st49 mutant phenotype. PHENOTYPE:
|
|
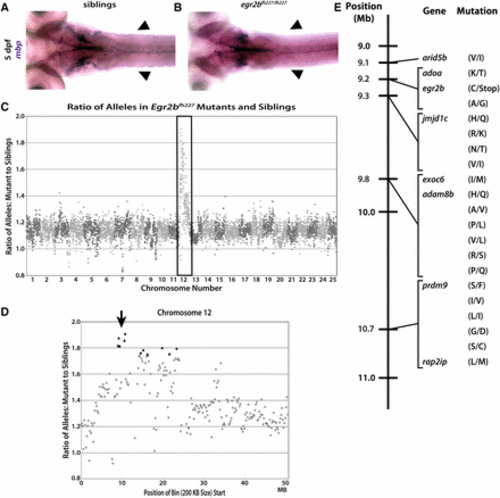
The fh227 allele is accurately mapped to chromosome 12 and egr2b using gDNA extracted from fixed tissue. Dorsal view of mbp expression in 5 dpf zebrafish using WISH and the mbp riboprobe shows phenotypically normal expression (purple) of mbp along the pLLNs (arrowheads) (A) compared to severely reduced mbp expression along the pLLN (arrowheads) in egr2bfh227/ fh227 mutants (B). (C) When the ratio of variant to reference alleles in the mutant pool is compared to the sibling pool and graphed across the whole genome for egr2bfh227, a clear spike on chromosome 12 is observed (box). This spike indicates genotypic linkage to the trait used to sort the mutant and sibling pools. (D) When looking at chromosome 12, egr2bfh227 is linked to a single region centered ∼10 Mb (arrow). (E) Linkage map of the egr2bfh227 allele showing the 21 different homozygous, nonsynonymous, protein-coding SNPs in the single chromosome 12 region linked to the decreased mbp expression in the PNS which was used to sort the mutant and sibling pools. The single introduced stop is the mutation responsible for the egr2bfh227 mutant phenotype. |
|
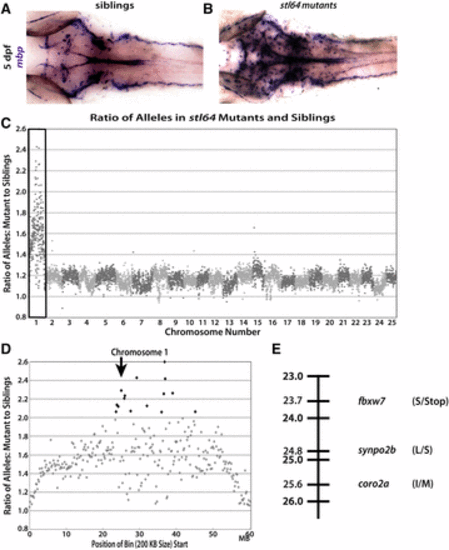
The stl64 phenotype is linked to chromosome 1 and is likely caused by a nonsense mutation in fbxw7. Dorsal views of 5 dpf zebrafish showing mbp expression by WISH using the mbp riboprobe. Normal expression in the CNS by phenotypically WT siblings (A) compared to the dramatic overexpression of mbp in the stl64 mutants (B) shows increased mbp expression in the stl64 mutants. (C) When the ratio of alleles in the mutant pool compared to the sibling pool is graphed across the whole genome for the stl64 allele, a clear spike on chromosome 1 (box) is observed for stl64. This spike indicates genomic linkage to the trait used to sort the mutant and sibling pools. (D) When viewing chromosome 1, stl64 is linked to a single region of the chromosome centered ∼24 Mb (arrow). (E) Linkage map of the stl64 allele showing the three protein-coding, homozygous, nonsynonymous SNP linked to the increased mbp expression in the CNS which was used to sort the mutant and sibling pools. The most deleterious SNP is the introduced stop codon in fbxw7. |
|
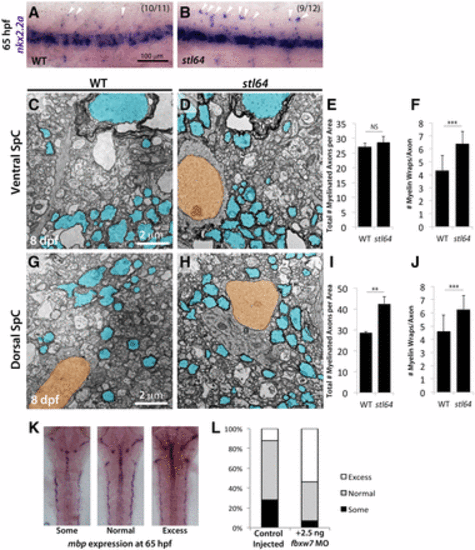
The fbxw7stl64 allele phenocopies the fbxw7vu56 allele. (A and B) stl64 mutants display more nkx2.2a+ cells in the dorsal spinal cord (SpC) relative to their WT siblings at 65 hpf. TEM analysis of the ventral (C–F) and dorsal (G–J) SpC at 8 dpf shows that fbxw7stl64 mutants have more myelinated axons in the dorsal SpC (I) and thicker myelin in both regions (F and J). (K and L) Transient morpholino (MO) knockdown of fbxw7 in WT embryos results in mbp overexpression at 65 hpf compared to control-injected siblings. Error bars are SD. ** P < 0.01, *** P < 0.001. NS, not significant. |