Fig. 2
- ID
- ZDB-FIG-240126-78
- Publication
- Cruz-Samperio et al., 2023 - Modular Bioorthogonal Lipid Nanoparticle Modification Platforms for Cardiac Homing
- Other Figures
- All Figure Page
- Back to All Figure Page
|
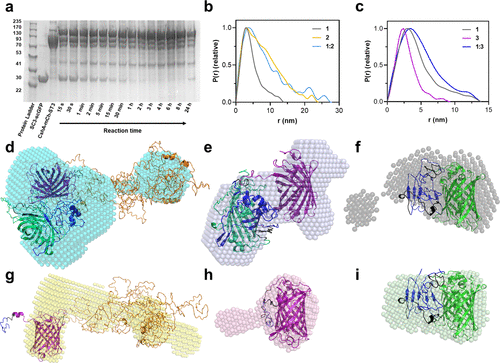
Characterization of reaction between protein constructs containing SpyCatcher and SpyTag. (a) SDS-PAGE monitoring of the reaction between 1 (10 μM [SC3-scGFP][S], ca. 42.5 kDa) and 2 (10 μM CshA-mCh-ST3, ca. 112 kDa) to yield 1:2 (CshA-mCh-ST3-[SC3-scGFP][S], ca. 155 kDa). (b) Pair-distance distribution function P(r) calculated using ScÅtter from synchrotron radiation small-angle X-ray scattering (SR-SAXS) data of 1 (shown in gray, χ2 = 1.396), 2 (shown in yellow, χ2 = 1.255), and the reaction product between both 1:2 (shown in light blue, χ2 = 1.292). (c) Pair-distance distribution function P(r) calculated using ScÅtter from synchrotron radiation small-angle X-ray scattering (SR-SAXS) data of 1, 3 (mCh-ST3, shown in purple, χ2 = 1.110), and the reaction product between both 1:3 (mCh-ST3-[SC3-scGFP][S], shown in navy blue, χ2 = 1.245). (d–h) Overlay of protein crystal structures and ab initio bead models computed from the SR-SXS data of (d) reaction product of CshA-mCh-ST3 and [SC3-scGFP][S] (1:2, bead model shown in cyan), (e) reaction product of mCh-ST3 and [SC3-scGFP][S] (1:3, bead model shown in navy blue), (f) [SC3-scGFP][S] (1, bead model shown in gray), (g) CshA-mCh-ST3 (2, bead model shown in orange), (h) mCh-ST3 (3, bead model shown in purple), and (i) SC3-scGFP (bead model shown in green). Proteins were modeled with I-TASSER, (32−34) and the models were selected depending on the known structures of SC3–ST3 (shown in navy blue), sfGFP (shown in green), and mCherry (shown in yellow). I-Tasser model C-scores: (d) −3.45, (e) −2.62, (f–i) −2.62, (g) −4.05, and (h) −0.62. |