Fig 1
- ID
- ZDB-FIG-200610-22
- Publication
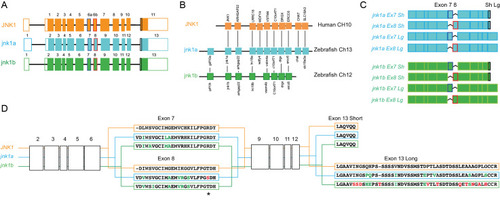
- Santos-Ledo et al., 2020 - Alternative splicing of jnk1a in zebrafish determines first heart field ventricular cardiomyocyte numbers through modulation of hand2 expression
- Other Figures
- All Figure Page
- Back to All Figure Page
|
( |