|
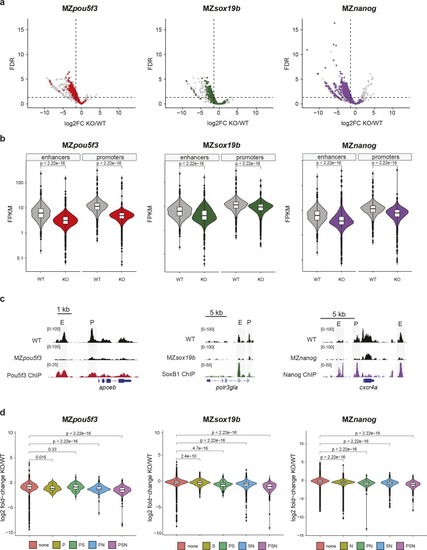
Pou5f3, Sox19b and Nanog regulate chromatin accessibility at gene regulatory elements.a) Volcano plots showing the log2 fold change in accessibility in MZpou5f3, MZsox19b and MZnanog mutants compared to wild-type embryos at oblong stage. Sites bound by Pou5f1, SoxB1, and Nanog are indicated in red, green and purple respectively. Non-bound sites are indicated in grey. b) Violin plots show the accessibility value (FPKM) at promoters and putative enhancers in wild-type embryos compared to the respective TF mutants. Significance of differences was tested using paired t-tests. c) Genome browser snapshots show accessibility tracks in MZpou5f3, MZsox19b and MZnanog mutants and wild-type embryos at oblong stage, for target genes of the respective TFs [15,25]. ChIP-seq tracks for the respective TFs at blastula stage are from data in [23,24]. d) Violin plots show the log2 fold change in mutants over wild-type at sites bound by different combinations of Pou5f3, SoxB1 and Nanog. p-values are shown for differences between binding sites as assessed by one-sided Wilcoxon tests.
|