- Title
-
Transcriptome analysis of atad3-null zebrafish embryos elucidates possible disease mechanisms
- Authors
- Ezer, S., Ronin, N., Yanovsky-Dagan, S., Rotem-Bamberger, S., Halstuk, O., Wexler, Y., Ben-Moshe, Z., Plaschkes, I., Benyamini, H., Saada, A., Inbal, A., Harel, T.
- Source
- Full text @ Orphanet J Rare Dis
|
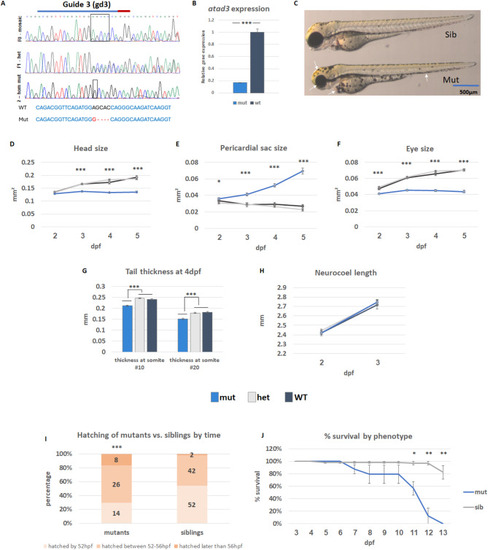
Characterization of |
|
Mitochondrial characterization and ATP content of |
|
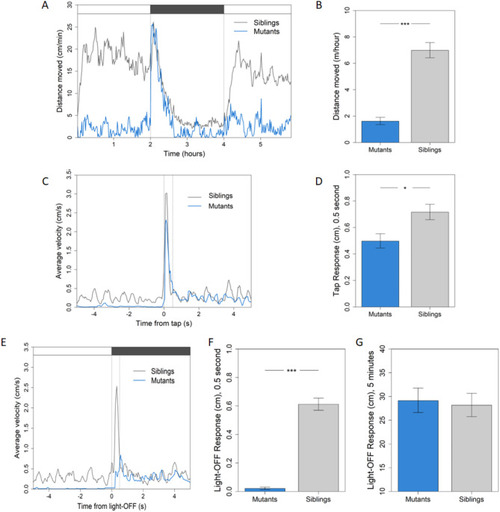
Locomotor activity of |
|
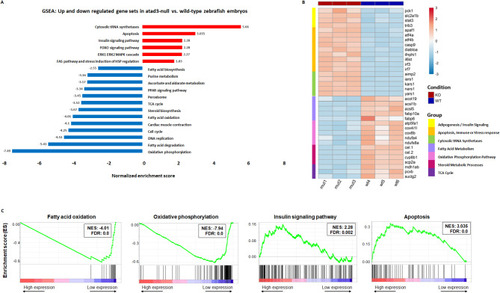
RNA-seq comparing mut 3 dpf embryos to their WT siblings. ( |
|
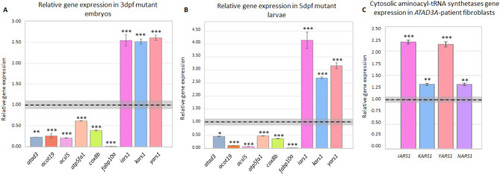
RT-qPCR validations for RNA-seq results. ( |