- Title
-
Identification of a PRPF4 Loss-of-Function Variant That Abrogates U4/U6.U5 Tri-snRNP Integration and Is Associated with Retinitis Pigmentosa
- Authors
- Linder, B., Hirmer, A., Gal, A., Rüther, K., Bolz, H.J., Winkler, C., Laggerbauer, B., Fischer, U.
- Source
- Full text @ PLoS One
|
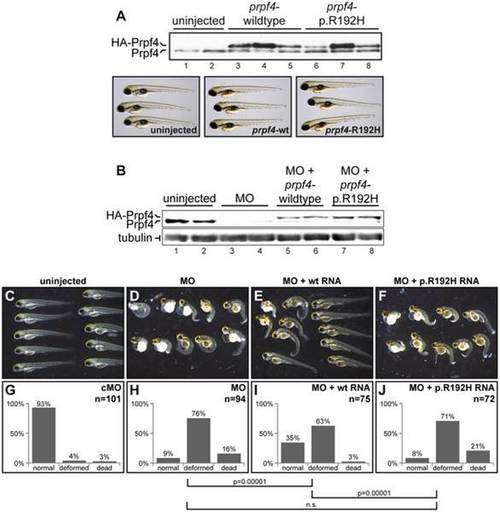
The p.R192H variant leads to a loss of function in vivo. (A) Expression of zebrafish Prpf4 carrying a mutation corresponding to p.R192H does not have a dominant negative effect. Upper panel: Western blot detection of endogenous Prpf4 and exogenous HA-tagged Prpf4 in embryos injected with RNAs indicated above. Lower panel: No defects in embryonic development were observed upon overexpression of wildtype or p.R192H Prpf4. (B) Western blot showing Prpf4 protein levels in zebrafish injected with the indicated combinations of prpf4 morpholino (MO) and RNA. (C–F) The p.R192H mutant fails to rescue a lethal prpf4 deficiency. Representative selections of 3 days post fertilization (dpf) larvae from embryos injected with the indicated combinations of prpf4 morpholino (MO) and RNA. (G–J) Quantification of normal, deformed and dead animals from four independent rescue experiments. The rescue effect (H vs. I) and the loss of function effect of the p.R192H mutation (I vs. J) were highly significant (Pearson X2 test; n is the total number of injected animals). Please note that the control experiments (uninjected, MO and MO + wt RNA), but not the characterization of p.R192H, were previously published as part of a larger knockdown study 22. As the characterization of p.R192H has been performed in the context of this study, we reproduce here the control data for clarity (with permission from Oxford University Press). PHENOTYPE:
|
|
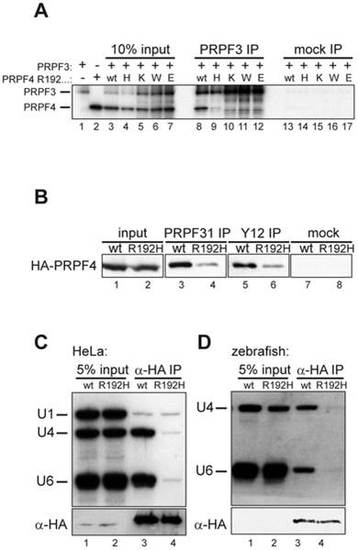
The arginine residue at position 192 is essential for integration of PRPF4 into snRNPs. (A) A series of mutants at position 192 was analyzed for binding to PRPF3. Neither lysine (positively charged) nor tryptophan (aromatic) nor glutamate (negatively charged) were tolerated at this position. (B) Reduced co-precipitation of p.R192H PRPF4 with snRNPs. HEK293 cells were transfected with wildtype and mutant HA-PRPF4 and lysates were immunoprecipitated using an anti-PRPF31 antibody (lanes 3 and 4), anti-Sm-antibody Y12 (lanes 5 and 6) or mock precipitated (lanes 7 and 8). Lanes 1 and 2 show 10% of the input used for the immunoprecipitations. (C) PRPF4 p.R192H fails to associate with U4 and U6 snRNA. HEK293 cells were transfected with HA-tagged PRPF4 in either wildtype or p.R192H mutant form. Anti-HA immunoprecipitation was controlled by Western blot (lower panel) and co-precipitated di-snRNP RNAs U4 and U6 were analyzed by Northern blotting (upper panel). U1 snRNA served as a specificity control. (D) snRNA co-immunoprecipitation experiments using zebrafish embryo lysates. Embryos were co-injected with 50 pg wildtype or p.R192H HA-prpf4 mRNA and 0.5 ng prpf4-MO to facilitate snRNP integration of exogenous protein. At 10 hpf, extracts were prepared and immunoprecipitations were analyzed as described in (C). |