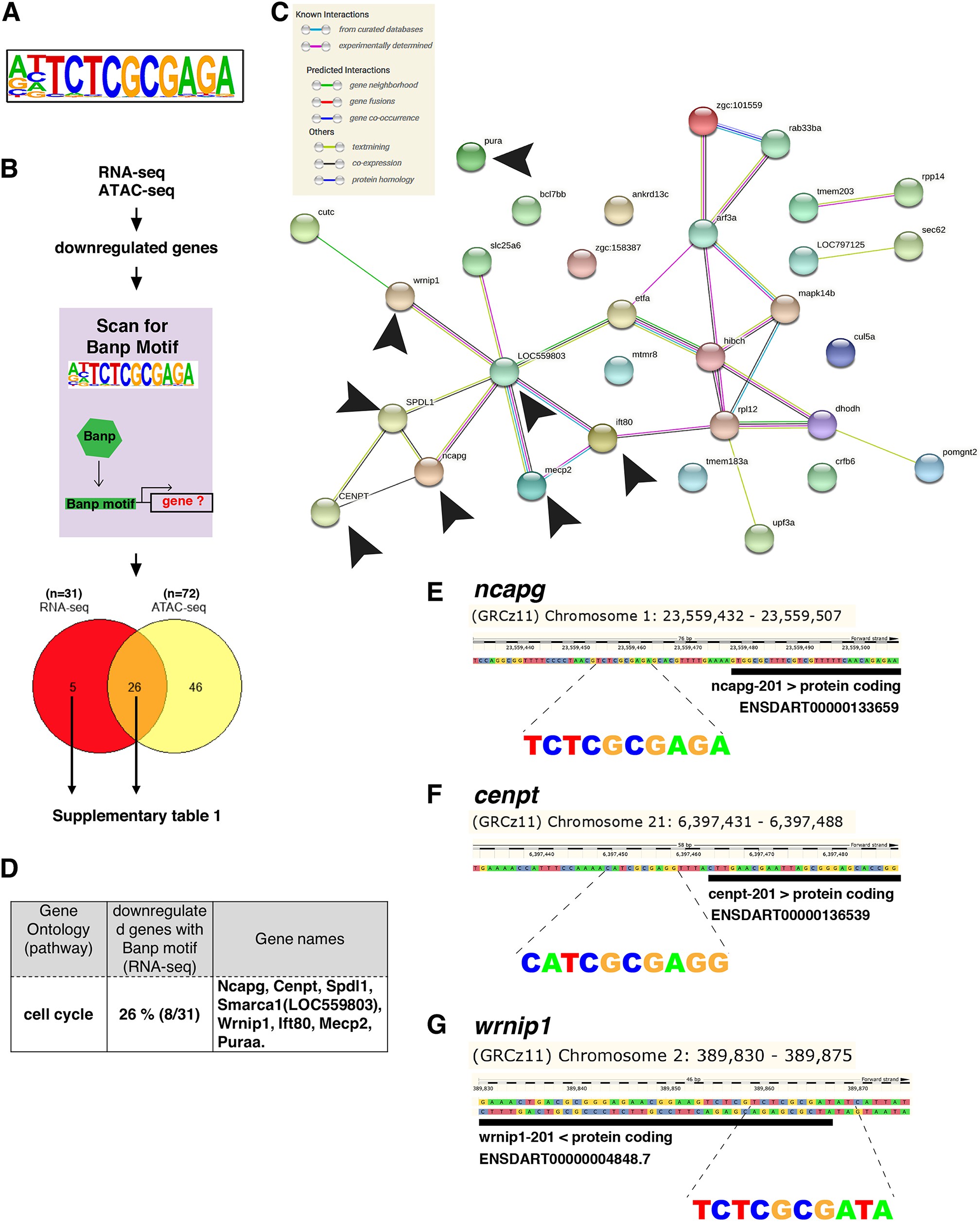
Fig. 6
(A) The motif enriched in genomic regions in which chromatin accessibility is reduced in banprw337 mutants. This sequence shows a high similarity to the Banp motif. (B) A schematic diagram representing the flow of RNA-sequencing combined with ATAC-sequencing to find candidates for zebrafish Banp direct-target genes. HOMER was used to scan for Banp motifs in downregulated candidate genes, from RNA- and ATAC-sequencing in banprw337 mutants. RNA sequencing yielded 31 candidates, whereas ATAC-sequencing yielded 72, with 26 target genes consistent in both. These 26 candidate genes are the most likely to be target genes of Banp. (C) STRING interactome analysis showing the interaction of Banp motif-containing genes downregulated in RNA-sequencing. Black arrowheads specify genes related to cell-cycle regulation. (D) Banp target candidate genes related to cell-cycle regulation. Gene ontology (GO) analysis was applied to 31 Banp motif-containing genes downregulated in RNA-sequencing. Twenty-six percent of these genes are engaged in the cell-cycle regulation. Others participate in a variety of non-categorized pathways. (E) Banp motif upstream to the 5’-UTR of ncapg. The black box represents the 5’-UTR. (F) Banp motif upstream to the 5’-UTR of cenpt. The black box represents the 5’-UTR. (G) Banp motif upstream to the 5’-UTR of wrnip1. The black box represents the 5’-UTR.
Banp promotes expression of genes containing Banp motifs.
Image
Figure Caption
Figure Data
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Elife