Figure 1
- ID
- ZDB-FIG-190723-701
- Publication
- Cioni et al., 2018 - Axon-Axon Interactions Regulate Topographic Optic Tract Sorting via CYFIP2-Dependent WAVE Complex Function
- Other Figures
- All Figure Page
- Back to All Figure Page
|
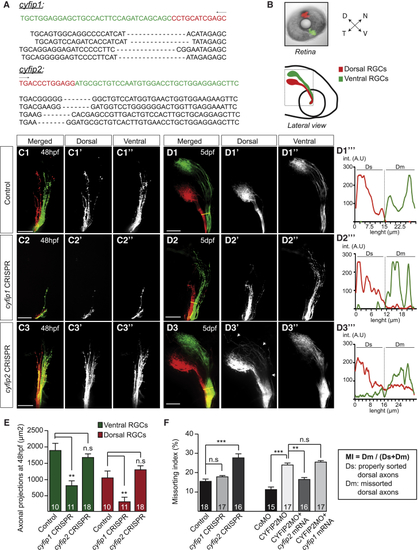
Differential Function of CYFIP1 and CYFIP2 during RGC Axonal Development (A) Sequence analysis of the whole zebrafish embryo injected with cas9 mRNA + cyfip1 or cyfip2 gRNAs. The target site is indicated on the sequence (green), followed by 4 examples of corresponding mutated regions. (B) DiI (red) and DiO (green) fluorescent dyes were injected in the zebrafish embryo retina at 5 dpf. The dashed line denotes the confocal imaging area of the optic tract (OT). (C1–D3′′′) Dorsal (D) (C1′–C3′, D1′–D3′) and Ventral (V) (C1′′–C3′′, D1′′–D3′′) RGC projections were analyzed in control embryos (cas9 mRNA + gRNA control) (C1, D1), cyfip1 CRISPR-injected embryos (cas9 mRNA + gRNA cyfip1) (C2, D2), and cyfip2CRISPR-injected embryos (cas9 mRNA + gRNA cyfip2) (C3, D3) at 48 hpf (C) and 5 dpf (D). Arrows in (D3′) show missorted dorsal axons in the OT. Yellow lines in (D1)–(D3) indicate the reference line used for quantification of the missorting index (MI). Examples of DiI (dorsal) and DiO signals plotted along the reference line corresponding to sorted (D1′′′, D2′′′) or misprojected (D3′′′) RGC axons (int., intensity; A.U, Arbitrary Units). (E) Quantifications of D and V axonal projection area in the OT at 48 hpf. (F) The missorting index (MI) was quantified as the ratio of the intensity signal of the missorted D (Dm) axons to all the D axons (Dm+Ds). For gRNA cyfip1-injected embryos, only the embryos showing an axon growth phenotype were quantified (n = 18 embryos). Error bars represent SEM. ∗∗p < 0.01, ∗∗∗p < 0.001, n.s., non significant (Mann-Whitney test for E and F). The number of zebrafish analyzed is indicated on the bars. Scale bars: 50 μm (C1–D3). See also Figure S1.
|
| Fish: | |
|---|---|
| Knockdown Reagents: | |
| Observed In: | |
| Stage Range: | Long-pec to Day 5 |