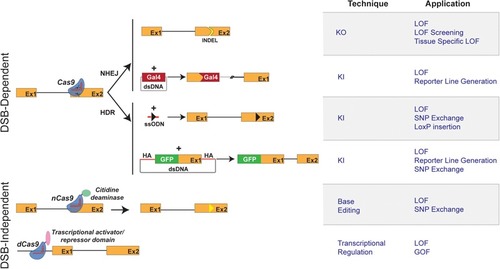
Schematic representation of CRISPR/Cas9 techniques and applications in zebrafish. (Left) Graphical representation of CRISPR/Cas9-based methods available for genome engineering in zebrafish. Double strand break (DSB)-Dependent techniques: the Cas9 endonuclease/sgRNA complex induces DSB at the target genomic locus. The NHEJ (non-homologous end joining) DNA repair mechanism leads to the appearance of insertion/deletion (INDELs) which can result in a shift of the open reading frame of the targeted genomic locus, thereby causing gene inactivation (knockout: KO). The targeted insertion (knockin: KI) of donor DNA sequences coding for reporter genes (e.g., Gal 4 transcriptional trans-activator) is, in most reports, mediated by NHEJ. The Homology Directed Repair mechanism allows the precise integration of exogenous DNA at a chosen genomic locus. Single-stranded oligodeoxynucleotide (ssODN) or Double stranded vectors harboring homology arms (HA) can be used as donor DNA for the KI of point mutations and reporter genes. Double strand break (DSB)-independent techniques: modified versions of the Cas9 protein, not displaying endonuclease activity, are used. The fusion of a cytidine deaminase to a Cas9 nickase (nCas9) allows a DSB-independent irreversible conversion of one targeted base to another (Base Editing and introduction of Single Nucleotide Polymorphisms, SNPs). Dead Cas9 (dCas9), when fused to a repressor domain or activator domain, can interact to either activate or repress transcription. (Right) Table describing the applications of CRISPR/Cas9-based methods depicted in the left panel.
|

