Fig. 1
- ID
- ZDB-FIG-150312-40
- Publication
- Irion et al., 2014 - Gap junctions composed of connexions 41.8 and 39.4 are essential for colour pattern formation in zebrafish
- Other Figures
- All Figure Page
- Back to All Figure Page
|
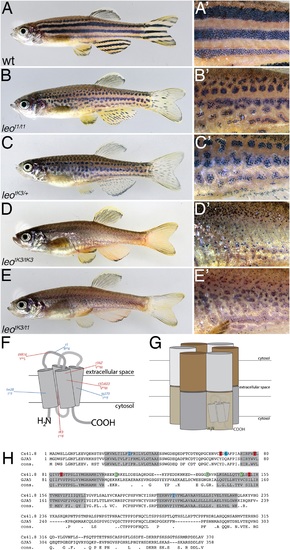
The leo mutant phenotype. Wild-type zebrafish (A) show a pattern of dark and light stripes on the body and on anal- and tail-fins. At higher magnification (A'), dark melanophores in the stripe regions and orange xanthophores in the light stripe regions are discernible. In mutants homozygous for leot1 (B) and heterozygous for leotK3 (C), the stripes are dissolved into spots. Clusters of melanophores are still visible (B' and C'). Fish homozygous for leotK3 (D) or trans-heterozygous for leotK3 over leot1 (E) show an identical phenotype of a completely dissolved pattern. Individual melanophores that hardly cluster together are still present, mostly associated with blue iridophores (D' and E'). In (F) a cartoon of Connexin 41.8 is depicted showing the positions of the leo mutations. Gap junctions are composed of two hemi-channels in adjacent cells. Each hemi-channel is made of six connexin subunits, they can be identical (homomeric) or different (heteromeric). In (G) a heteromeric/heterotypic gap junction is schematically shown. An alignment of the amino acid sequence from zebrafish Cx41.8 with its human orthologue, GJA5, is shown in (H). The transmembrane regions are shaded in grey. Newly identified point mutations in Connexin 41.8 are highlighted in red: leotNR16: Y66S; leotNZ: V79M; leot3OJ022: V85M; leotK3: I152F. Already known alleles are highlighted in blue: leotw28: I31F; leot1: R68X; leotq270: I202F. Polymorphisms found in sequences from wild-type fish are highlighted in green: 106 R/K, 136 G/R; 149 V/I. |
| Fish: | |
|---|---|
| Observed In: | |
| Stage: | Adult |