- Title
-
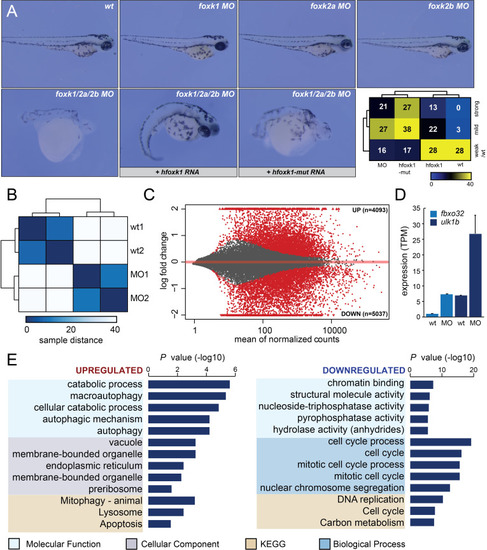
Depletion of Foxk transcription factors causes genome-wide transcriptional misregulation and developmental arrest in zebrafish embryos
- Authors
- Geng, F.S., de la Calle-Mustienes, E., Gómez-Skarmeta, J.L., Lister, R., Bogdanovic, O.
- Source
- Full text @ MicroPubl Biol
|
EXPRESSION / LABELING:
PHENOTYPE:
|