- Title
-
Mitochondrial ClpX Activates a Key Enzyme for Heme Biosynthesis and Erythropoiesis
- Authors
- Kardon, J.R., Yien, Y.Y., Huston, N.C., Branco, D.S., Hildick-Smith, G.J., Rhee, K.Y., Paw, B.H., Baker, T.A.
- Source
- Full text @ Cell
|
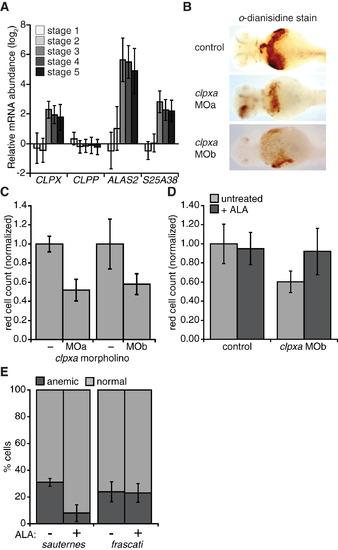
mtClpX Is Important for Vertebrate Heme Biosynthesis and Erythropoiesis (A) Relative mRNA abundance for human CLPX, CLPP, ALAS2, and SLC25A38 (indicated as S25A38) throughout erythropoiesis, as indicated in a microarray dataset described in Novershtern et al. (2011). Erythroid development stages were defined by cell-type-specific markers as follows: 1, CD34+ CD71+ GlyA 2, CD34 CD71+ GlyA 3, CD34 CD71+ GlyA+; 4, CD34 CD71low GlyA+; 5, CD34 CD71 GlyA+. (B) o-dianisidine staining (brown) for hemoglobinized red cells in zebrafish embyos. Embyros were grown from zygotes injected at the one- to two-cell stage with clpxa-targeting morpholinos (MOa and MOb) or uninjected zygotes (control). (C) Erythrocyte development at 72 hpf was quantified by flow cytometry, using dissociated cells from Tg(globin-LCR:eGFP) zebrafish. p d 0.01 for erythrocyte reduction by clpxa knockdown with either morpholino. (D) Rescue of clpxa MOb-induced anemia by ALA supplementation. Tg(globin-LCR:eGFP) zebrafish embryos were supplemented with 2 mM ALA from 24 to 72 hpf, upon which GFP+ erythrocytes were quantified by flow cytometry. p = 0.025 for rescue of anemia in clpxa knockdown embryos by ALA supplementation. (E) Heterozygous sauternes (ALAS2 mutant) or frascati (SLC25A37 mutant) zebrafish were crossed, and progeny were grown for 72 hpf, with or without ALA supplementation as in (D). Anemia was assayed by o-dianisidine staining. p = 0.04 for rescue of anemia in sauternes+/ progeny by ALA. n = 52 for sauternes ALA; n = 43 for sauternes +ALA; n = 98 for frascati ALA; n = 122 for frascati +ALA. Error bars represent mean ± SD. See also Figure S5. PHENOTYPE:
|
|
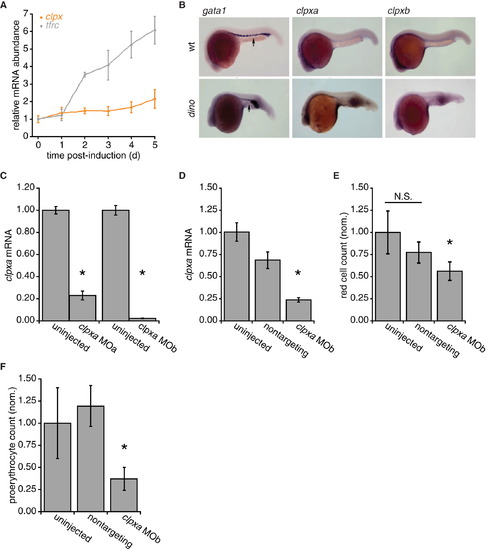
mtClpX in Vertebrate Heme Biosynthesis and Erythropoiesis, Related to Figure 6 (A) Relative mRNA abundance for mouse clpx and tfrc (transferrin receptor; included as a positive control for genes strongly induced during erythropoiesis) throughout erythropoiesis. mRNA was collected from MEL cells at indicated times after induction of erythropoiesis by the addition of 1.5% DMSO to growth medium. mRNA was quantified by qPCR and normalized to a control mRNA (hprt). p < 0.05 for upregulation of clpx at days 2, 4, and 5, and for tfrc at days 2–5. (B) In situ hybridization to determine expression pattern of ClpX isoforms in D. rerio embryos. gata-1 mRNA was used to delineate the ventral blood island (intermediate cell mass, ICM, arrows) in wt embryos. Dino mutants, which are ventralized due to a defect in chordin, exhibit expansion of the erythropoietic marker gata-1. D. rerio clpxa and clpxb exhibit nearly ubiquitous expression, with higher expression observed in neural tissues 24 hr post-fertilization. (C) D. rerio clpxa morpholino-injected embryos have reduced clpxa mRNA levels. clpxa mRNA levels in uninjected and clpxa morpholino-injected D. rerio embryos at 72 hpf are displayed (as detected by qRT-PCR, normalized to hprt control). p < 0.001. (D) clpxa mRNA levels in uninjected, nontargeting morpholino-injected, and clpxa morpholino-injected D. rerio embryos at 24 hpf are displayed. p < 0.01, ANOVA. (E) Erythrocyte development at 72 hr post-fertiliztion (hpf) was quantified by flow cytometry, using dissociated cells from Tg(globin-LCR:eGFP) zebrafish, in which all erythrocytes are GFP-labeled. p < 0.01, ANOVA. (F) FACS analysis of uninjected, nontargeting morpholino-injected and clpx MOb-injected Tg(gata1:GFP) zebrafish embryos at 24hpf. clpx morphants display a reduction in GFP+ erythroid progenitor cells. p < 0.0001, ANOVA. Error bars represent mean ± SD. EXPRESSION / LABELING:
PHENOTYPE:
|
Reprinted from Cell, 161, Kardon, J.R., Yien, Y.Y., Huston, N.C., Branco, D.S., Hildick-Smith, G.J., Rhee, K.Y., Paw, B.H., Baker, T.A., Mitochondrial ClpX Activates a Key Enzyme for Heme Biosynthesis and Erythropoiesis, 858-867, Copyright (2015) with permission from Elsevier. Full text @ Cell