- Title
-
In toto imaging of the migrating Zebrafish lateral line primordium at single cell resolution
- Authors
- Nogare, D.D., Nikaido, M., Somers, K., Head, J., Piotrowski, T., Chitnis, A.
- Source
- Full text @ Dev. Biol.
|
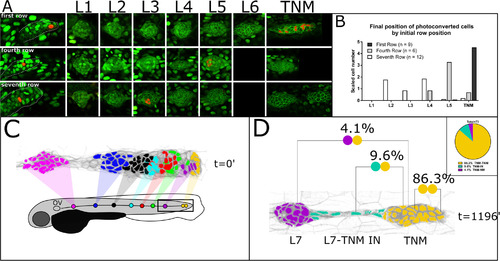
Spatial pattern of cell fate the lateral line primordium. A. Photoconversion of single cells in the first, fourth and seventh row of the PLLp at 22hpf (left-most panel), and subsequent position of cells in the formed lateral line at 48hpf (right panels). B. Quantification of cell fate from photoconversion data in A. C Fate of cells in PLLp at L1 deposition color coded by neuromast number. Uncolored cells are either Different Fate (DF) cells or interneuromast cells. D Division classes in lineages contributing to the terminal neuromast cluster (yellow). The proportion of each indicated division is shown in the inset. Only a small percentage of lineages contribute to both the terminal cluster and to a more trailing neuromast (in this case, L7). |
|
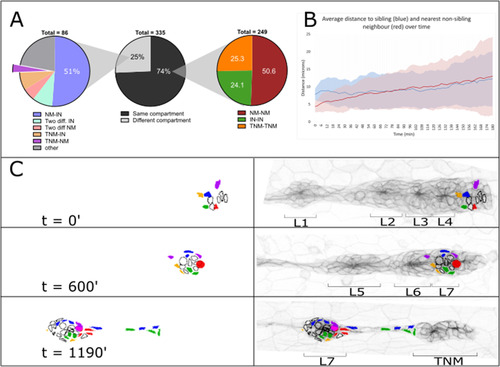
Quantification of cell fate in the PLLp. A Pie charts of cell divisions classified by daughter cell fate. Center panel: Divisions classified by whether both daughters contribute to the same (black) or different (grey) structures in the LL. Right panel: same-fate divisions classified by compartment (NM, IN or TNM). Left panel: different-fate divisions classified by eventual fate of daughter cells. Expanded wedge represents divisions where one daughter cell remains in the terminal cluster and another is deposited in a non-terminal neuromast. “Other” represents divisions in which daughters themselves undergo divisions. B Plot of average separation distance following division over time between sibling cells (blue) and between dividing cell and nearest non-neighbor sibling. Shaded regions represent one standard deviation. C Frames from movie reconstruction showing all cell lineages that will contribute to the L7 neuromast. Black outlined cells represent lineages in which all daughters will contribute exclusively to L7, colored lineages represented three lineages which will contribute to L7 as well as additional compartments (primarily the L7-TNM interneuromast cell population). |
|
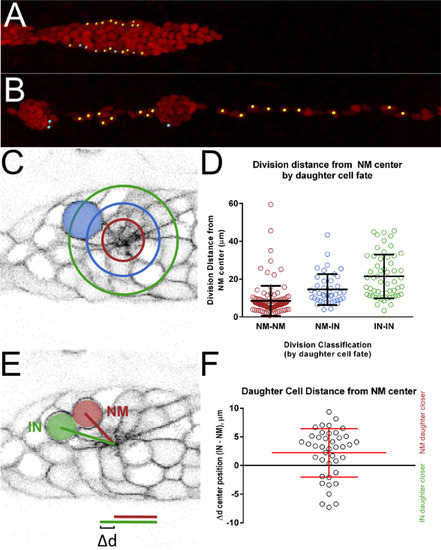
Position of dividing cells by distance from the center of neuromasts. A, B frames from timelapse movie showing position of future interneuromast cells within the PLLp. Yellow dots mark interneuromast cells, blue dots mark peripheral support cells within the neuromast. Only one movie out of three that were tracked is shown. C Average radial distance from center of neuromast of divisions classified by division type (red NM-NM, green IN-IN and blue NM-IN. D Quantification of radial distance of division from neuromasts center by cell fate. p<0.0001, one-way ANOVA. E Schematic of distance from center of neuromast for NM and IN cells immediately after a NM-IN division. Δd is calculated by the difference in the distance between IN and NM cells. F Plot of Δd for all cells. Positive values indicate NM-fated daughter cell closer to NM center, negative values indicate IN-fated daughter cell closer to NM center. |
Reprinted from Developmental Biology, 422(1), Nogare, D.D., Nikaido, M., Somers, K., Head, J., Piotrowski, T., Chitnis, A., In toto imaging of the migrating Zebrafish lateral line primordium at single cell resolution, 14-23, Copyright (2017) with permission from Elsevier. Full text @ Dev. Biol.