- Title
-
Accurate Reconstruction of Cell and Particle Tracks from 3D Live Imaging Data
- Authors
- Liepe, J., Sim, A., Weavers, H., Ward, L., Martin, P., Stumpf, M.P.
- Source
- Full text @ Cell Syst
|
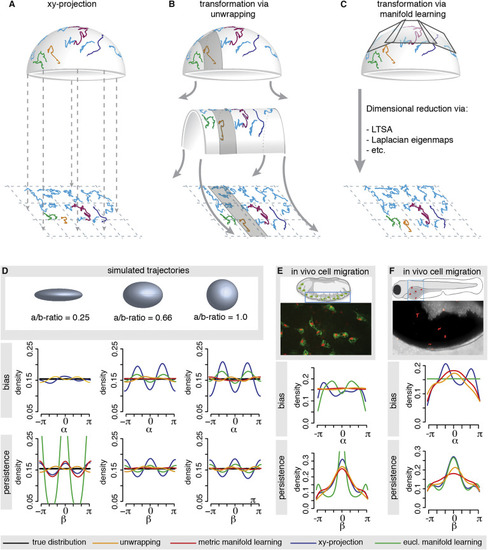
Methods for Manifold Learning and Applications (A-C) Shown are example trajectories on a hemi-sphere and their transformation via one of the discussed methods. The 3D cell tracks are simply projected onto the xy-plane (A). The 3D cell tracks are transformed via unwrapping (B) or via well-known manifold learning methods (e.g., LTSA) (C). (D) Random walk trajectories (in absence of any bias or persistence) are simulated on the displayed curved surfaces and then transformed with xy-projection, unwrapping, Euclidian manifold learning and Riemannian, or metric, manifold learning, respectively. The resulting bias and persistence distributions are compared with the respective true distribution (black), which are uniform for this random walk model. For the ellipsoid with the most extreme aspect ratio (i.e., the “thinnest” shape), the manifold learning approach was unsuccessful as it incorrectly interpreted the data as belonging to a 1D line. In this case, there was insufficient data to correctly reveal the spatial extent of one dimension. (E) Application of the unwrapping method and manifold learning methods to haemocyte cell tracks extracted from a D. melanogaster embryo and their comparison to the xy-projection. Shown is a schematic of the embryo and a snapshot from the video microscopy imaging. Haemocytes (green) were tracked via their nucleus (red). (F) Application of the unwrapping method and manifold learning methods to neutrophil cell tracks extracted from the epidermis overlying the yolk syncytium of a zebrafish and their comparison to the xy-projection. The epidermis was wounded with a laser before image acquisition. Shown is a schematic of the zebrafish with the imaged area and a snapshot from the video microscopy imaging with the neutrophils in red. |