- Title
-
Regulation of β1 Integrin-Klf2-Mediated Angiogenesis by CCM Proteins
- Authors
- Renz, M., Otten, C., Faurobert, E., Rudolph, F., Zhu, Y., Boulday, G., Duchene, J., Mickoleit, M., Dietrich, A.C., Ramspacher, C., Steed, E., Manet-Dupé, S., Benz, A., Hassel, D., Vermot, J., Huisken, J., Tournier-Lasserve, E., Felbor, U., Sure, U., Albiges-Rizo, C., Abdelilah-Seyfried, S.
- Source
- Full text @ Dev. Cell
|
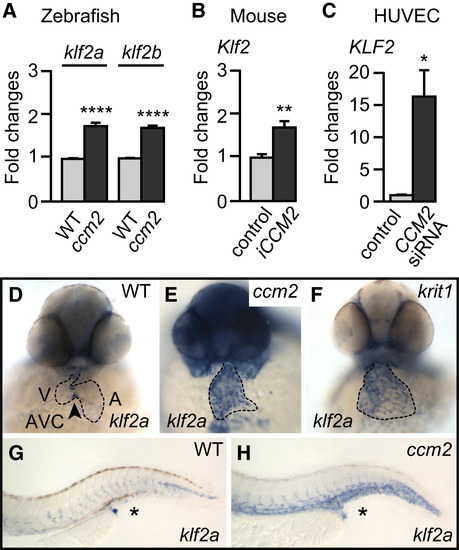
Loss of CCM Proteins Causes Elevated Levels of klf2 Expression (A–C) Loss of Ccm2 causes elevated levels of klf2a and klf2b mRNA in zebrafish at 48 hrpf (A), of Klf2 mRNA in mouse cerebellar tissue of iCCM2 mice at P15 (B), and of KLF2 mRNA in HUVECs (C), as determined by qRT-PCR. (D–H) At 48 hrpf, whole-mount in situ hybridizations show that klf2a is misexpressed within the entire endocardium of krit1ty219c and ccm2m201 mutants (D–F) and is elevated within arteries and veins in the tail region (asterisk) of ccm2m201 mutants (H) compared to WT embryos (G). Statistical data are means and SEM; p < 0.05; p < 0.01; p < 0.001; p < 0.0001; see Supplemental Experimental Procedures for details about statistical data analysis. EXPRESSION / LABELING:
|
|
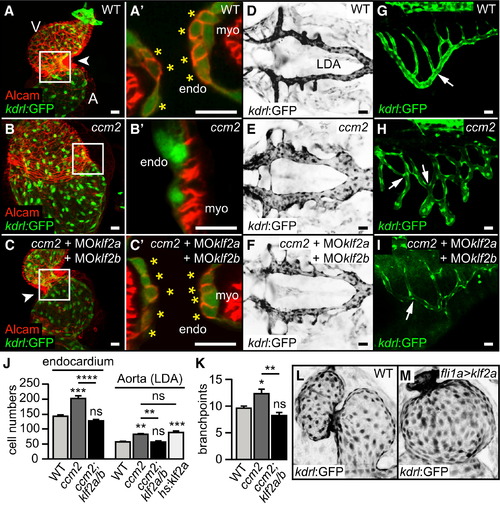
Elevated Levels of klf2 mRNA Cause Morphogenetic Defects in Zebrafish ccm Mutants (A–I) Rescue of cardiac malformations, LDA defects, and SIV defects in zebrafish ccm2m201 mutants (B, B′, E, and H) by targeting klf2a/b with MOs (C, C′, F, and I). (A–C) Shown are hearts of different genotypes at 48 hrpf or (A′, B′, and C′) details with single confocal plane sections of the atrioventricular canal (AVC) region (arrowheads, white box). Endocardial cushion cells are marked by Tg(kdrl:GFP)s843 and Alcam staining (asterisks). (D–F) Dorsal views of the LDA marked by Tg(kdrl:GFP)s843 (inverted images) in different genotypes at 48 hrpf. (G–I) Side view of the SIV marked by Tg(kdrl:GFP)s843 in different genotypes at 72 hrpf. Vessel branch point defects (arrows) within the SIV are reduced by targeting klf2a/b in ccm2m201 mutants (H and I). (J) Endocardial and LDA cell numbers are elevated in ccm2m201 mutants at 48 hrpf, as well as upon overexpression of klf2a from a heat-shock-inducible promoter. (K) Quantifications of the number of branch points of the SIV in different genotypes at 72 hrpf. (L and M) Endothelial overexpression of klf2a in Tg(fli1ep:Gal4FF)ubs2; Tg(UAS:klf2a)ig1–3; Tg(kdrl:GFP)s843 transgenic embryos is sufficient to phenocopy the ccm2m201 mutant cardiac ballooning phenotype. Endo, endocardium; myo, myocardium; A, atrium; V, ventricle. Scale bars, 25 µm. Statistical data are means and SEM; ns, not significant; p < 0.05; p < 0.01; p < 0.001; p < 0.0001; see Supplemental Experimental Procedures for details about statistical data analysis. EXPRESSION / LABELING:
PHENOTYPE:
|
|
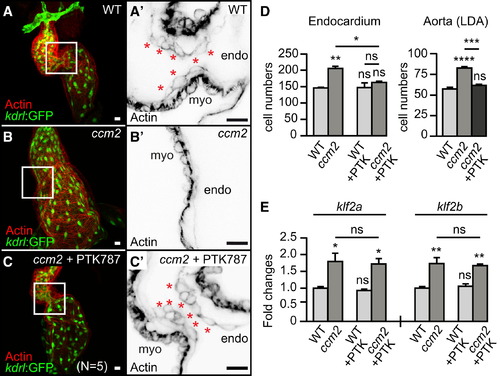
Enhanced VEGFR Signaling Is Involved in the Zebrafish ccm2m201 Mutant Cardiac Phenotype (A–C) Rescue of cardiac malformations in ccm2m201 mutants at 48 hrpf after treatment with the VEGFR inhibitor PTK787 between 15 and 17 hrpf. (A–C) Shown are hearts marked by endocardial Tg(kdrl:GFP)s843 and actin staining at 48 hrpf or (A′, B′, and C′) details with single confocal plane sections of the AVC region (white box in A, B, and C). Endocardial cushion cells are characterized by their cuboidal shape (asterisks) (n = 5/5 genotyped ccm2m201 mutants completely rescued by PTK787 treatment). (D) Treatment of ccm2m201 mutants with PTK787 between 15 and 17 hrpf significantly reduces the endocardial overproliferation phenotype, and treatment between 24 and 48 hrpf suppresses the overgrowth of the LDA. (E) The phenotypic rescue of ccm2m201 mutants by treatment with PTK787 between 15 and 48 hrpf does not affect the elevated expression levels of klf2a and klf2b as detected by qRT-PCR. Endo, endocardium; myo, myocardium. Scale bars, 25 µm. Statistical data are means and SEM; ns, not significant; p < 0.05; p < 0.01; p < 0.001; p < 0.0001; see Supplemental Experimental Procedures for details about statistical data analysis. |
|
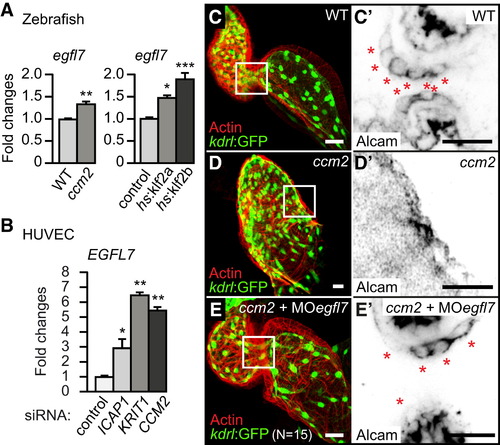
Elevated Expression Levels of egfl7 mRNA Are Involved in the ccm2m201 Mutant Phenotype (A) Loss of Ccm2 or overexpression of klf2a/b in Tg(hsp70l:klf2a)md8 or Tg(hsp70l:klf2b)md9 transgenic embryos causes elevated levels of egfl7 mRNA, as determined by qRT-PCR. (B) Similarly, EGFL7 mRNA is upregulated in HUVECs upon knockdown of ICAP1, KRIT1, or CCM2 by siRNA treatment. (C-E′) Cardiac morphology and cardiac cushion formation are rescued by egfl7 MO injection (n = 15/15 genotyped ccm2m201 mutants completely rescued by egfl7 MO). Shown are hearts marked by endocardial Tg(kdrl:GFP)s843 and actin staining at 48 hrpf or (C′-E′) details with single confocal plane sections of the AVC region. Endocardial cushion cells are marked by the expression of Alcam (inverted image) (asterisks). Scale bars, 25 µm. Statistical data are means and SEM; p < 0.05; p < 0.01; p < 0.001; see Supplemental Experimental Procedures for details about statistical data analysis. |
|
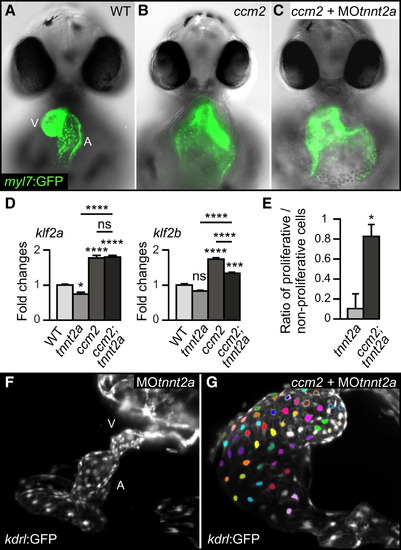
Blood-Flow-Independent Expression of klf2a/b mRNA in ccm2m201 Mutants (A–C) The ccm2m201 mutant cardiac ballooning phenotype is flow independent. Frontal views of 48 hrpf hearts of the myocardial reporter line Tg(myl7:GFP)twu34 (Huang et al., 2003) show that cardiac expansion is not rescued in ccm2m201 mutant; tnnt2a morphants that lack blood flow. (D) The expression levels of klf2a and klf2b mRNA, as measured by qRT-PCR, are elevated in ccm2m201 mutant; tnnt2a morphants that lack blood flow. (E–G) Loss of Ccm2 results in overproliferation of the endocardium under no-flow conditions. (E) Quantifications of the ratio of proliferating versus nonproliferating endocardial cells between 33 and 53 hrpf in tnnt2a morphants compared with ccm2m201 mutant;tnnt2a morphants based on SPIM recordings (n = 2). (F) The endocardium of tnnt2a morphants is not proliferative as indicated by the lack of color-coded proliferative cells (still image derived from Movie S1, which is based on SPIM time-lapse recordings of endocardial morphogenesis marked by Tg(kdrl:GFP)s843 between 33 and53 hrpf). (G) The combined loss of Ccm2 and Tnnt2a is characterized by a high rate of proliferative events (indicated by uniquely colored daughter cells derived from proliferative events; still image derived from Movie S2). A, atrium; V, ventricle. Statistical data are means and SEM; ns, not significant; p < 0.05; p < 0.01; p < 0.001; p < 0.0001; see Supplemental Experimental Procedures for details about statistical data analysis. |
|
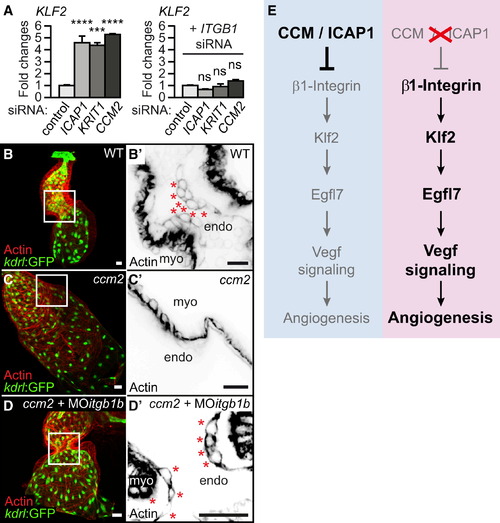
Misexpression of KLF2 Depends on Integrin β1 (A) qRT-PCR shows that elevated expression of KLF2 mRNA in ICAP1-, KRIT1-, or CCM2-silenced HUVECs is partly suppressed upon simultaneously interfering with β1 integrin (ITGB1). (B–D) Morpholino knockdown of zebrafish integrinβ1b completely restores cardiac morphology (marked by Tg(kdrl:GFP)s843 and actin staining) and cardiac cushion formation in zebrafish ccm2m201 mutants at 48 hrpf. (B′–D′) Shown are single confocal plane sections with details of the AVC region (white box in B–D) with endocardial cushion cells as marked by actin (asterisks). (E) Model of CCM/ICAP1-controlled proangiogenic activity of the β1 integrin-Klf2 mechanotransduction pathway. Endo, endocardium; myo, myocardium. Scale bars, 25 µm. Statistical data are means and SEM; ns, not significant;p < 0.001; p < 0.0001; see Supplemental Experimental Procedures for details about statistical data analysis. |
|
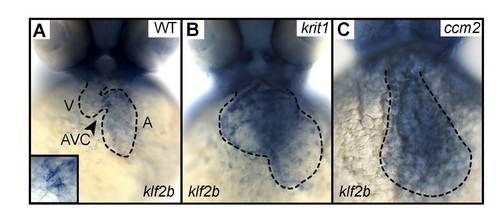
Endocardial expression of klf2b mRNA is misregulated in zebrafish ccm mutants. (A) Whole-mount in situ hybridization reveals that klf2b is expressed at the atrioventricular canal (AVC) at 48 hpf in WT. (B,C) Loss of Krit1 or of Ccm2 causes elevated expression of klf2b throughout the entire ballooning endocardium.EXPRESSION / LABELING:
|
Reprinted from Developmental Cell, 32, Renz, M., Otten, C., Faurobert, E., Rudolph, F., Zhu, Y., Boulday, G., Duchene, J., Mickoleit, M., Dietrich, A.C., Ramspacher, C., Steed, E., Manet-Dupé, S., Benz, A., Hassel, D., Vermot, J., Huisken, J., Tournier-Lasserve, E., Felbor, U., Sure, U., Albiges-Rizo, C., Abdelilah-Seyfried, S., Regulation of β1 Integrin-Klf2-Mediated Angiogenesis by CCM Proteins, 181-190, Copyright (2015) with permission from Elsevier. Full text @ Dev. Cell