- Title
-
Xirp proteins mark injured skeletal muscle in zebrafish
- Authors
- Otten, C., van der Ven, P.F., Lewrenz, I., Paul, S., Steinhagen, A., Busch-Nentwich, E., Eichhorst, J., Wiesner, B., Stemple, D., Strähle, U., Fürst, D.O., and Abdelilah-Seyfried, S.
- Source
- Full text @ PLoS One
|
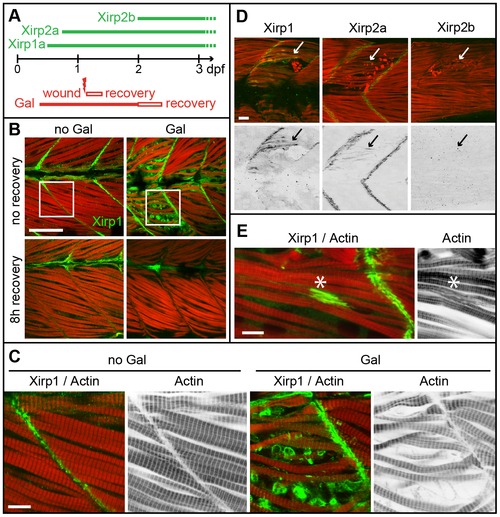
Expression and localization of Xirps within Galanthamine- and laser-induced myocellular wounds. (A) Schematic diagram summarizing the temporal order of Xirp expression within somitic muscle and in myocellular wounding assays. (B) Galanthamine (GAL) treatment between 80% epiboly and 2 dpf causes severe disruptions of somitic muscle organization and myofibrillar disarray (red: Actin) in 2 dpf zebrafish embryos. Notably, Xirp1 (green) is strongly expressed and localizes within cells most strongly disrupted by the treatment. These effects are completely reversible within several hours of recovery. Scale bar: 50 μm. (C) Details from inserts indicated in B (green: Xirp1; red: Actin). Scale bar: 10 μm. (D, E) Similarly, laser-induced muscle injury induces ectopic Xirp1 and Xirp2a localization to damaged myofibrils. In comparison, Xirp2b is not yet expressed at 33 hpf. Green: Xirp1, Xirp2a or Xirp2b; red: Actin. Arrows indicate the position of laser-induced injury within somitic tissue. Scale bars: 10 μm. Embryos were injured at 27 hpf (D) or 29 hpf (E) and fixed at 33 hpf (D) or 31.5 hpf (E), respectively. |
|
Structure of the zebrafish Xirp family and gene expression patterns. (A) Phylogenetic analysis of Xirp family members. The zebrafish Xirp family comprises three members which are orthologous to their two mammalian counterparts. Phylogenetical distances calculated according to the ClustalW slow/accurate method are indicated. Danio rerio (D.r.); Gallus gallus (G.g.); Mus musculus (M.m.). (B) As for mammalian Xirp1, intraexonic splicing of zebrafish xirp1 results in three alternative isoforms: xirp1α (6894 bp), xirp1β (4452 bp), xirp1γ (2006 bp). Xirp1β is identical to the N-terminus of Xirp1α and harbours Xin-repeats; Xirp1γ is identical to the C-terminus of Xirp1α and harbours a Filamin C binding domain (FilC BD). Antibodies specific against isoforms Xirp1α/β (P43) and Xirp1α/γ (P47) were generated. Black asterisks indicate the position of protein truncation in the xirp1sa0046 mutant. Concerning Xirp2a and Xirp2b, differential splicing leads to various isoforms classified as long (containing Xin-repeats) or short (lacking a large exon encoding Xin-repeats and resulting in the encoding of a LIM domain). Here, we represented schematically the long isoforms and their respective antibody. (C) Summary panel of xirp gene family expression with whole-mount in situ hybridization probes designed to exclusively cover the long isoforms of these genes. xirp1 has the earliest and widest expression of all three xirp genes. After initial expression within adaxial cells during somitogenesis, xirp1 expression decreases within somitic tissue after 24 hpf. xirp1 is the only xirp family member harboring Xin-repeats that is expressed within the heart during cardiogenesis. In comparison, xirp2a and xirp2b mRNAs are strongly expressed within somitic muscle after 24 hpf where they localize to the somite boundaries. Both xirp1 and xirp2a are expressed within other types of skeletal muscle including those of the fin buds, whereas xirp2b expression remains restricted to somitic muscle. Asterisks indicate the heart; arrowheads point at fin buds. Black scale bars: 250 μm, red scale bars: 200 μm. EXPRESSION / LABELING:
|
|
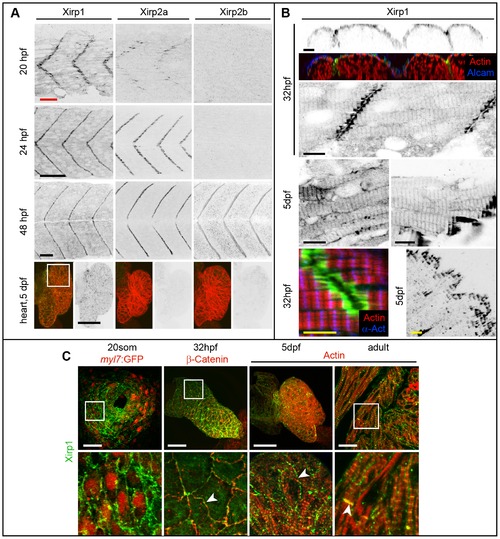
Expression and localization of Xirp proteins during skeletal and cardiac muscle development in WT. (A) Xirp expression patterns overlap within skeletal muscle between 20–48 hpf. In comparison, only Xirp1 is expressed within the cardiac tissue at 5 dpf. Green: Xirp1, Xirp2a or Xirp2b; red: Actin. Red scale bar: 25 μm, black scale bars: 50 μm. (B) Top panel shows cross section planes of confocal z-scan projections through somitic tissue. Note that Xirp1 (green) is most strongly expressed within the outer layer of slow muscle cells, as marked by Alcam (blue) expression (red: Actin). Whereas most of Xirp1 localizes in a staircase-like pattern to the MTJ, some protein is also detectable in a repetitive pattern at the z-discs of myofibrils and at the lateral plasma membrane, depending on the tissue. Different skeletal muscle types are shown at 5 dpf, whereas somitic muscle is shown at 32 hpf. Green: Xirp1; red: Actin; blue:α- Actinin. Black scale bars: 10 μm, yellow scale bars: 5 μm. (C) Expression and localization of Xirp1 during cardiogenesis and in adult cardiac tissue. Bottom row represents higher magnification images (from inserts, if there is a white box on the top picture) with details on Xirp1 subcellular localization at the ICD of cardiomyocytes (arrowheads). Green: Xirp1; red: myl7:GFP, β-Catenin or Actin. Scale bars: 50 μm. EXPRESSION / LABELING:
|
|
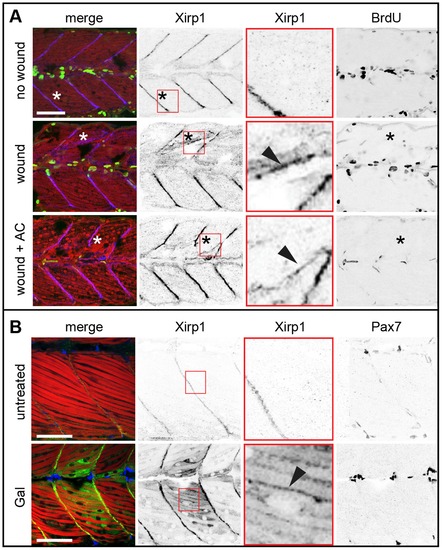
Xirp1 marks wounded skeletal muscle cells prior to de novo cell proliferation. (A) Laser-induced myocellular injuries in 33 hpf old zebrafish embryos after BrdU pulse labeling (between 24–33 hpf). Embryos were wounded at 31 hpf and left to recover for 2 hours. Xirp1+ tissue is devoid of BrdU+ proliferative cells. Treatment of laser-induced myocellular injuries with the proliferation inhibitor Aphidicolin (together with BrdU between 24–33 hpf) does not affect expression and localization of Xirp1 within damaged tissue. The efficacy of the Aphidicolin treatment is evident from strongly reduced BrdU labelling. Asterisks indicate the position of laser-induced injury within damaged tissue. Red inserts show Xirp1 localization within damaged tissue. Green: BrDU; red: Actin; blue: Xirp1. (B) Consistent with the lack of proliferating cells within Xirp1+ damaged tissue, the distribution of Pax7+ external cells is not changed compared to control conditions. Arrowheads mark ectopic Xirp1. Red inserts show Xirp1 localization within damaged tissue. Green: Xirp1; red: Actin; blue: Pax7. All scale bars: 50 μm. |
|
Clonal analysis of laser-induced myocellular injuries. (A–B) Laser-injury and recovery within the same embryo. (A) Left: DIC image superimposed with image of individual muscle cells marked by Tg[βactin:α-actinin-gfp] expression (false colored in yellow). Right: laser-injury was performed by targeting one of the α-Actinin-GFP positive muscle cells. Immediately upon laser-injury, α-Actinin-GFP expression within the damaged cell is diminished (arrowheads). Scale bar: 50 μm. (B) Confocal images of two z-scan planes of an immunohistochemical staining show strong Xirp1 expression within 2.5 hours after laser injury in muscle cells directly adjacent to the targeted cell that was most severely affected by the laser-injury (arrowheads). Pictures on the right are details from the insert (white box on the bottom left picture). Green: βactin:α-Actinin-GFP; red: Actin; blue: Xirp1. Scale bar: 50 μm. (C) Confocal z-scan projection of an immunohistochemical staining 5 hours after laser-induced injury reveals that strong Xirp1 expression correlates with a pattern of Tropomyosin distribution that is different from unaffected regions of the somite. Green: Tropomyosin; red: Actin; blue: Xirp1. Scale bar: 20 μm. |
|
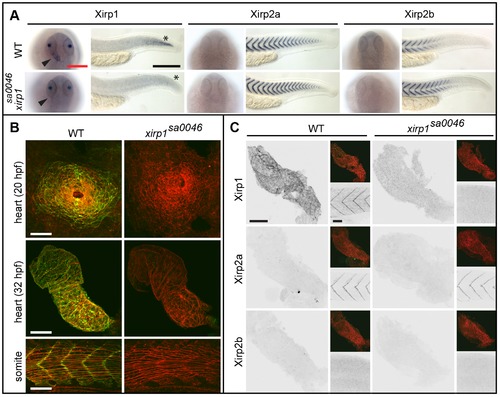
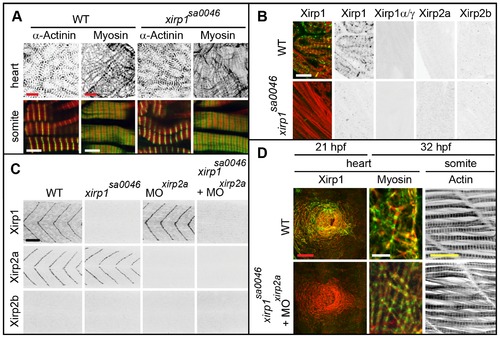
Normal cardiogenesis and skeletal muscle development in the xirp1sa0046 mutant. (A) Whole-mount in situ hybridization of xirp1, xirp2a and xirp2b on 24 hpf WT and xirp1sa0046 mutants reveal a loss of xirp1 expression in heart and muscle of xirp1sa0046 mutants. In contrast, neither xirp2a nor xirp2b expression are affected by lack of xirp1. In particular, neither xirp2a nor xirp2b are upregulated in the hearts of xirp1sa0046 mutants. Arrowheads indicate the position of the heart tube. Asterisks indicate the tip of the tail. Red scale bar: 200 μm, black scale bar: 250 μm. (B) Lack of xirp1 mRNA expression corresponds with the complete absence of Xirp1 (green) within the 20 hpf heart cone, 32 hpf heart tube, and 21 hpf skeletal muscle. The Actin counterstaining (red) demonstrates that cardiogenesis and skeletal muscle organization at these stages is comparable between WT and xirp1sa0046 mutants. Scale bars: 50 μm. (C) Immunohistochemical stainings for Xirp1, Xirp2a and Xirp2b on 24 hpf WT and xirp1sa0046 mutant hearts show complete absence of Xirp2a and Xirp2b. Top inserts depict overlay images of Actin (red) and Xirp1, Xirp2a or Xirp2b (green). Within the WT somitic muscle, both Xirp1 and Xirp2a are expressed whereas Xirp1 is absent in xirp1sa0046 mutant somitic tissue (bottom inserts). Scale bars: 50 μm. EXPRESSION / LABELING:
PHENOTYPE:
|
|
Normal cardiogenesis and myofibrillogenesis in the absence of all Xirps. (A) Neither cardiac nor skeletal muscle sarcomeric organization of myofibrils is affected in 48 hpf xirp1sa0046 mutants. Green: α-Actinin or Myosin; red: Actin. Red scale bars: 10 μm, white scale bars: 5 μm. (B) Sectioned adult cardiac tissue reveals that neither Xirp2a nor Xirp2b are expressed to compensate for the loss of Xirp1. Also, the xirp1sa0046 mutant lacks all three Xirp1 isoforms. Therefore, Xirps are not required for development or maintenance of cardiac tissue. Green: Xirp1; red: Actin. Scale bar: 10 μm. (C) Complete absence of all Xirps within skeletal muscle prior to 24 hpf in xirp1sa0046/xirp2aMO mutant/morphants. Localization of Xirp1 is not affected in xirp2a morphants and, conversely, Xirp2a localization is normal in xirp1sa0046 mutants. Scale bar: 50 μm. (D) Complete loss of Xirps in xirp1sa0046/xirp2aMO mutant/morphants does not impair early cardiogenesis (at 21 hpf) or myofibrillogenesis (at 32 hpf). Green: Xirp1 or Myosin; red: Actin. Red scale bar: 50 μm, white scale bar: 5 μm, yellow scale bar: 20 μm. EXPRESSION / LABELING:
PHENOTYPE:
|
|
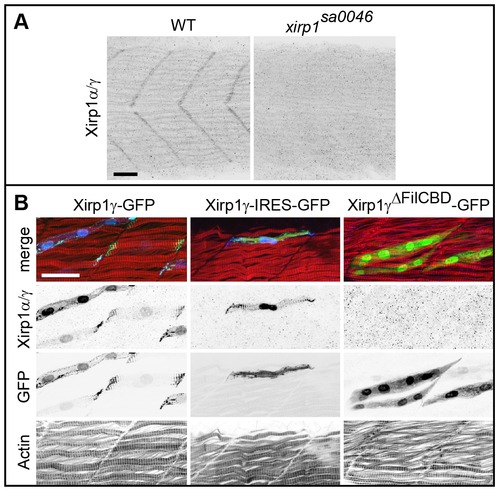
Characterization of the P47 antibody. (A) Myotendinous junction labeling by the P47 antibody which recognizes Xirp1α/γ is absent in xirp1sa0046 mutant embryos at 24 hpf. This staining was performed upon mild fixation. (B) Clonally expressed Xirp1γ is sensitively detected upon standard fixation conditions by the P47 antibody within somitic muscle tissue. Truncated Xirp1γΔFilCBD-GFP which lacks the Filamin C binding domain including the P47 epitope is not detected by the antibody. Under standard fixation conditions, Xirp1α/γ is not detected at the myotendinous junctions. Green: GFP or Xirp1γ-GFP fusion protein; red: Actin; blue: Xirp1α/γ. All scale bars: 25 μm. EXPRESSION / LABELING:
|
|
Efficient Morpholino antisense oligonucleotide-mediated knock-down of xirp2a does not affect myofibrillogenesis. (A) Phenotypically, xirp2a morphants are indistinguishable from WT embryos at 24 hpf. Scale bar: 250 μm. (B) Efficient gene knock-down of xirp2a is assessed by immunohistochemistry at 24 hpf. Green: Xirp2a; red: Actin. Scale bar: 20 μm. (C) Complete loss of Xirp2a does not affect myofibrillogenesis and correct sarcomeric organization of skeletal muscle as determined by immunohistochemistry using an antibody against the A/I junction epitope of Titin and rhodamine phalloidin to label sarcomeric Actin at 24 hpf. Green: Titin; red: Actin. Scale bar: 5 μm. PHENOTYPE:
|
|
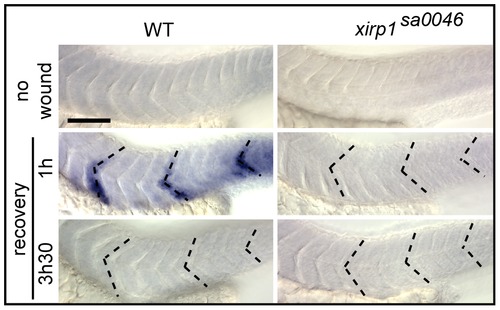
Rapid xirp1 transcriptional response to myocellular injury. Whole-mount in situ hybridizations on 33 hpf embryos which were laser-injured either at 29 hpf (bottom row) or at 32 hpf (middle row) at the level of three different somites (dotted lines). xirp1 mRNA transcriptional response occurs within one hour after injury and is no longer detectable at 3.5 hours after injury in WT. There is a lack of xirp1 mRNA expression upon laser-induced myocellular injury in xirp1sa0046 mutants. Scale bar: 100 μm. EXPRESSION / LABELING:
|
|
Sharacterization of interactions between Xirp1 and Filamin C or Enah/Vasp family members. (A) Table summarizing the results of direct yeast two hybrid experiments with the C-terminus of zebrafish Xirp1 (residues 2097–2297) and the GST-tagged domains 19–21 of zebrafish Filamin Ca (FlnCa) and b (FlnCb). The Filamin CbΔ (FlnCbΔ) isoform represents a splice variant lacking the unique insertion within domain 20 that is present in all Filamin C proteins. (B) Western blot overlay experiments show specific binding of the GST-tagged domains 19–21 of FlnCa and FlnCb to the blotted C-terminus of Xirp1. GST and FlnCbΔ do not bind, confirming that the unique insertion within domain 20 of Filamin Cb is necessary for the interaction. (C) Co-immunoprecipitation experiments confirm these interactions: FlnCa and FlnCb are co-precipitated, whereas GST alone is not. (D) Western blot overlay experiments show specific binding of T7-tagged N-terminus of zebrafish Xirp1 (residues 1–58) containing proline-rich repeat1 (PR1) to the blotted EEF-tagged EVH1 domains of zebrafish Enah, VaspI and VaspII. (E) Summary of binding assays shows the isoforms of Xirp1 identified in our study, and the respective binding sites for EVH1 domains of Enah and Vasp (PR1) and Filamin C (carboxy-terminus). Binding to F-actin is presumably mediated by Xin-repeats (red boxes). PRs are depicted by blue boxes. |