- Title
-
Caveolin-1 regulates dorsoventral patterning through direct interaction with beta-catenin in zebrafish
- Authors
- Mo, S., Wang, L., Li, Q., Li, J., Li, Y., Thannickal, V.J., and Cui. Z.
- Source
- Full text @ Dev. Biol.
|
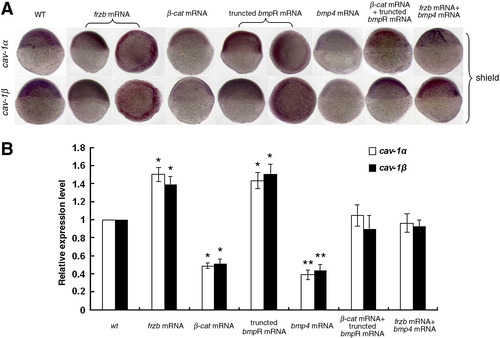
Transcriptional regulation of zebrafish cav-1 expression. Transcriptional expression of cav-1α and -1β was detected with whole-mount in situ hybridization (A) and real-time RT-PCR (B). Embryos at one-cell stage were injected with 300 pg frzb mRNA, 300 pg β-catenin 1 (β-cat) mRNA, 300 pg truncated bmpR mRNA, 300 pg bmp4 mRNA, 150 pg frzb mRNA plus 150 pg bmp4 mRNA, or 150 pg β-cat mRNA plus 150 pg truncated bmpR mRNA. Wild-type (WT) embryos were used as the control. Embryos at shield stage are shown in lateral views with the animal pole toward the top and the dorsal toward the right. Embryos injected with frzb mRNA and truncated bmpR mRNA were also oriented for an animal pole view with the dorsal toward the right. Real-time RT-PCR data represent mean ± S.D. from three independent experiments. *indicates p < 0.05 vs. wt and * *indicates p < 0.01 vs. wt. EXPRESSION / LABELING:
|
|
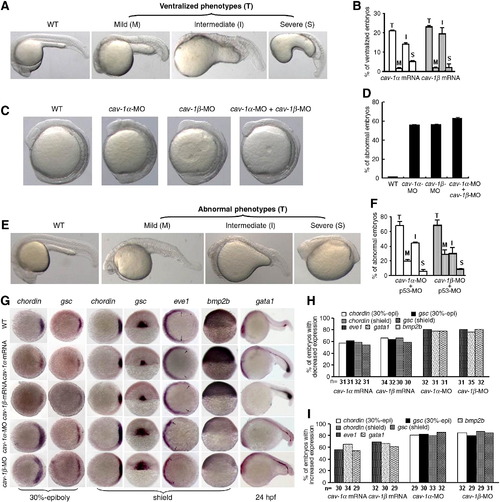
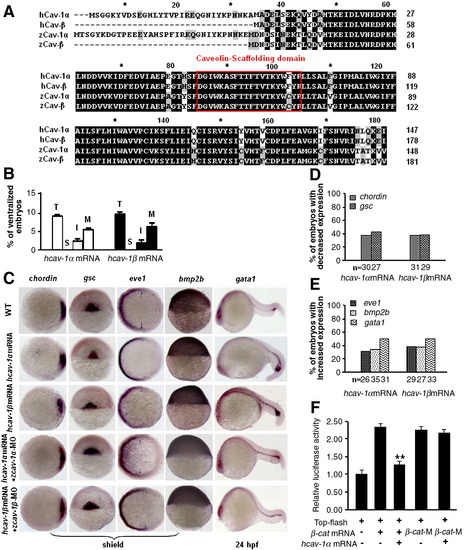
Effects of cav-1 overexpression and knockdown on zebrafish dorsoventral patterning. (A) Overexpression of cav-1 ventralizes zebrafish embryos. Embryos at one-cell stage were injected with 300 pg cav-1α or -1β mRNA. Wild-type (WT) embryos was used as the control. Total ventralized embryos (T) were divided into three groups: mild (M), intermediate (I) and severe (S). (B) Percentage of ventralized embryos for each group in (A). Data represent mean ± S.D. from three independent experiments. (C) Knockdown of cav-1 led to weakly dorsalized embryos with early tailbud protrusion at 12 hpf. Embryos at one-cell stage were injected with 10 ng cav-1α-MO, 10 ng cav-1β-MO, or 5 ng cav-1α-MO plus 5 ng cav-1β-MO. (D) Percentage of abnormal embryos at 12 hpf for each group in (C). (E) Knockdown of cav-1 led to abnormal embryos at 24 hpf. Embryos at one-cell stage were injected with 1 ng p53 MO plus 10 ng cav-1α-MO or -1β-MO. Total number of abnormal embryos (T) at 24 hpf was divided into three groups: mild (M), intermediate (I) and severe (S). (F) Percentage of abnormal embryos at 24 hpf for each group in (E). Data represent mean ± S.D. from three independent experiments. (G) Effects of cav-1 overexpression and knockdown on expression patterns of dorsoventral marker genes (indicated on the top). Developmental stages are shown at the bottom. Lateral views of embryos at shield stage with the dorsal toward the right for bmp2b. Animal pole views of embryos at shield stage with the dorsal toward the right for chordin, eve1 and gsc at 30% epiboly stage. Dorsal views of embryos at shield stage with the animal pole toward the top for gsc at shield stage. Lateral views of embryos at 24 hpf with the anterior to the left for gata1. Arrows point to the margins of expressed eve1. (H–I) Total number of embryos examined (indicated at the bottom) and percentage of embryos with decreased or increased expression of dorsoventral marker genes in (G). EXPRESSION / LABELING:
PHENOTYPE:
|
|
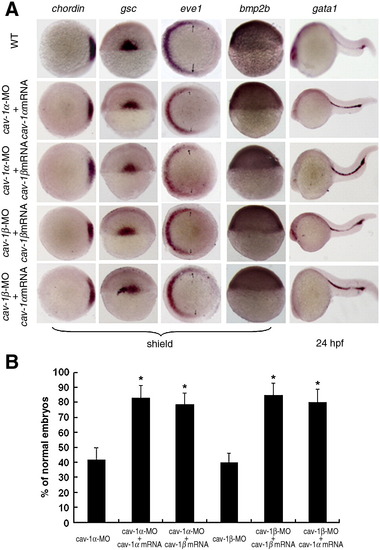
Abnormal phenotypes of Cav-1 knockdown embryos can be rescued by its capped mRNAs. (A) Abnormal expression patterns of dorsoventral marker genes caused by injection of Cav-1-MO can be rescued by its corresponding or another isoform mRNAs. Developmental stages are shown at the bottom. Animal pole views of embryos at shield stage with the dorsal toward the right for chordin and eve1 and arrows point to the margins of expressed eve1. Lateral views of embryos at shield stage with the dorsal toward the right for bmp2b. Dorsal views of embryos at shield stage with the animal pole toward the top for gsc. Lateral views of embryos at 24 hpf with the anterior to the left for gata1. (B) Percentage of normal embryos injected with 5 ng cav-1α-MO, 5 ng cav-1α-MO plus 300 pg cav-1α mRNA, 5 ng cav-1α-MO plus 300 pg cav-1β mRNA, 5 ng cav-1β-MO, 5 ng cav-1β-MO plus 300 pg cav-1β mRNA, 5 ng cav-1β-MO plus 300 pg cav-1α mRNA. Data represent mean ± S.D. from three independent experiments. *indicates p < 0.05 vs. cav-1α-MO or cav-1β-MO. |
|
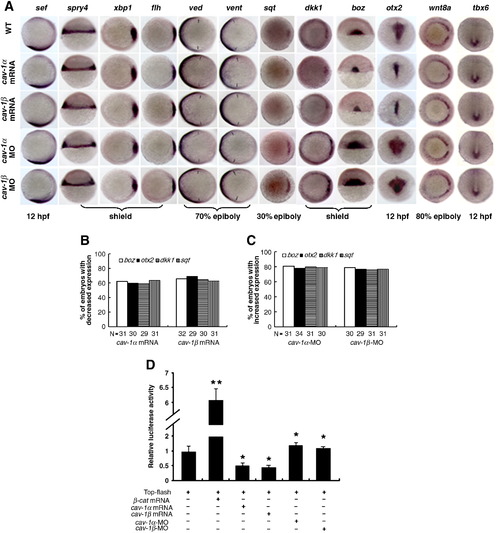
Cav-1 regulates zebrafish dorsoventral patterning via maternal Wnt signaling. (A) Expression patterns of reporter genes (indicated on the top) for FGF, Nodal, BMP and Wnt signalings in wild-type (WT) or embryos injected with 300 pg cav-1α mRNA, 300 pg cav-1β mRNA, 10 ng cav-1α-MO or 10 ng cav-1β-MO. Developmental stages are shown at the bottom. Lateral views of embryos at 12 hpf with anterior toward the top for sef. Dorsal views of embryos at shield stage with animal pole toward the top for spry4 and boz. Animal pole views of embryos at shield stage with the dorsal toward the right for xbp1, flh and dkk1. Animal pole views of embryos at 30% epiboly stage with the dorsal toward the right for squint (sqt). Animal pole views of embryos at 70% epiboly stage with the dorsal toward the right for ved and vent. Arrows point to the margins of expressed ved and vent. Dorsoanterior views of embryos at 12 hpf with the anterior toward the top for otx2. Vegetal views of embryos at 80% epiboly stage with dorsal toward the right for wnt8a. Dorsal views of embryos at 12 hpf with the anterior toward the top for tbx6. (B) Total number of embryos examined (indicated at the bottom) and percentage of embryos with decreased expression of genes sqt, dkk1, boz and otx2 in (A) after injection of cav-1 mRNA. (C) Total number of embryos examined (indicated at the bottom) and percentage of embryos with increased expression of genes sqt, dkk1, boz and otx2 in (A) after injection of cav-1-MOs. (D) Luciferase activity of Top-flash reporter for Wnt signaling was inhibited by injection of 300 pg cav-1α or -1β mRNA, but enhanced by injection of 10 ng cav-1α-MO or 10 ng cav-1β-MO. Injection of 50 pg mRNA for constitutively active β-catenin-1 (β-cat mRNA) stimulated activity of Top-flash reporter and was used as the positive control. Data for luciferase assays represent mean ± S.D. from three independent experiments. *indicates p < 0.05 and **indicates p < 0.01 vs. Top-flash-injected embryos. EXPRESSION / LABELING:
PHENOTYPE:
|
|
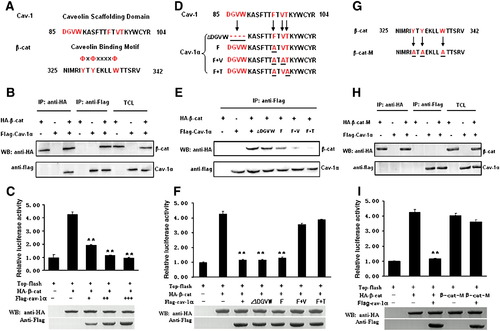
Biochemical analysis of the Cav-1 and β-catenin interaction. The following assays were performed in ZF4 cells. Immunoprecipitation assays (IP) and Western blots (WB) were performed to detect interaction between Flag-Cav-1 (or its Flag-mutants) and HA-β-cat (or HA-β-cat-M). Data for luciferase activity assays represent mean ± S.D. from three independent experiments. **indicates p < 0.01 vs. cells transfected with Top-flash plus β-cat. TCL indicates total cell lysates. (A) Critical amino acid residues in Cav-1 scaffolding domain and in putative Cav-1 binding motif of β-catenin were highlighted in red. (B) Reciprocal interaction between Flag-Cav-1 and HA-β-cat. (C) Luciferase activity of Top-flash reporter was inhibited in a dose-dependent manner by co-transfection of 50 ng β-cat plasmid DNA and 100 ng Top-flash reporter DNA with Cav-1α plasmid DNA at 25 ng (+), 50 ng (++) and 75 ng (+++). WB analysis showed the proper expression of β-cat and the Cav-1 expression in a dose-dependent manner. (D) Amino acid residues deleted or substituted in Cav-1 scaffolding domain were underlined. (E) Mutations in Cav-1 affect reciprocal interaction between Flag-Cav-1 and HA-β-cat. (F) Activation of Top-flash reporter by transfection of 50 ng β-cat plasmid DNA was inhibited by co-transfection of 75 ng plasmid DNA for Cav-1α, Cav-1αΔDGVW or Cav-1α/F, but not by co-transfection of 75 ng plasmid DNA for Cav-1α/F + V or Cav-1α/F + T. WB analysis showed the proper expression of β-cat and Cav-1. (G) Amino acid residues substituted in putative Cav-1 binding motif of β-catenin were underlined. (H) Substitution of three amino acid residues in putative Cav-1 binding motif of β-catenin disrupted the interaction between HA-β-cat and Flag-Cav-1. (I) Activation of Top-flash reporter by transfection of 50 ng β-cat-M plasmid DNA was not inhibited by co-transfection of 75 ng Cav-1α plasmid DNA. WB analysis showed the proper expression of β-cat and Cav-1. |
|
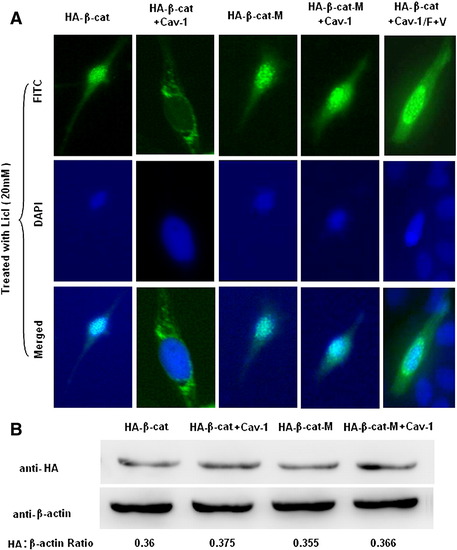
Overexpression of Cav-1 blocks the nuclear translocation of active β-catenin. (A) ZF4 cells on 35 mm dishes were transfected with 600 ng of each plasmid expressing HA-β-cat plus pRC/CMV2, HA-β-cat plus Cav-1, HA-β-cat-M plus pRC/CMV2, HA-β-cat-M plus Cav-1, or HA-β-cat plus Cav-1/F + T. Transfected cells were then treated with 20 mM LiCl for 24 h. Cellular distribution of HA-β-cat or HA-β-cat-M was shown by location of FITC signal. Nuclei were stained with DAPI. (B) Effects of Cav-1 on degradation of β-catenin were detected by Western immunoblotting. ZF4 cells on 35 mm dishes were transfected with 600 ng of each plasmid expressing HA-β-cat plus pRC/CMV2, HA-β-cat plus Cav-1, HA-β-cat-M plus pRC/CMV2, or HA-β-cat-M plus Cav-1. Expression of β-actin was used as loading control. |
|
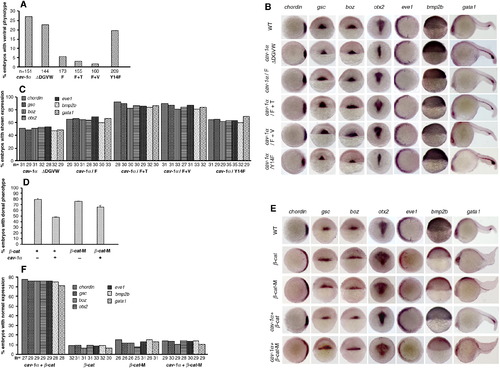
Genetic interaction between Cav-1 and β-catenin in zebrafish embryos. (A) Percentage of ventralized embryos after injection of 300 pg capped mRNA for Cav-1α or its mutants at one-cell stage. (B) Expression patterns of dorsoventral genes in embryos injected with 300 pg capped mRNA for Cav-1α or its mutants at one-cell stage. Developmental stages are shown at the bottom. Animal pole views of embryos at shield stage with the dorsal toward the right for chordin and eve1. Arrows point to the margins of expressed eve1. Dorsal views of embryos at shield stage with the animal pole toward the top for gsc and boz. Lateral views of embryos at shield stage with the dorsal toward the right for bmp2b. Dorsoposterior views of embryos at 12 hpf with the posterior toward the top for otx2. Lateral views of embryos at 24 hpf with the anterior to the left for gata1. (C) Statistical data for (B). Total numbers of embryos detected are indicated at the bottom. (D) Percentage of dorsalized embryos caused by injection of 150 pg β-cat mRNA plus 150 pg GFP mRNA, 150 pg β-cat mRNA plus 150 pg cav-1 mRNA, 150 pg β-cat-M mRNA plus 150 pg GFP mRNA, or 150 pg β-cat-M mRNA plus 150 pg cav-1 mRNA. (E) Expression patterns of dorsoventral genes in embryos from (D). (F) Percentage of normal embryos in (E) that were rescued by injection of 150 pg mRNA for cav-1α or its mutants. Total number of embryos examined was indicated at the bottom. |
|
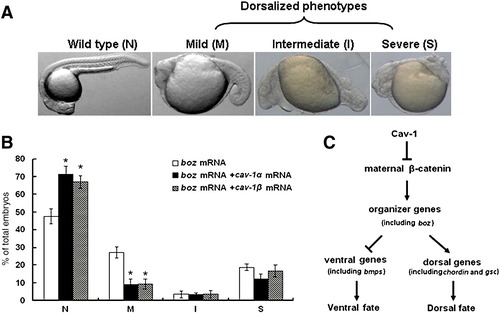
Ventralizing activity of Cav-1 is mediated by inhibition of Boz activity. (A) Overexpression of boz dorsalizes zebrafish embryos. Embryos at one-cell stage were injected with 5 pg boz mRNA. Wild-type embryos were used as the control (N). Dorsalized embryos were divided into three groups: mild (M), intermediate (I) and severe (S). (B) Percentage of normal embryos for each group. Embryos at one-cell stage were injected with 5 pg boz mRNA, 5 pg boz mRNA plus 150 pg mRNA for cav-1α, or 5 pg boz mRNA plus 150 pg mRNA for cav-1β. Data represent mean ± S.D. from three independent experiments. (C) A proposed model for regulation of dorsoventral patterning by Cav-1. Cav-1 inhibits activity of maternal β-catenin by limiting its nuclear translocation. Activation of Wnt/β-catenin signaling in the dorsal organizer promotes boz expression that inhibits expression of ventral genes, including bmp2b and bmp4, while stimulating expression of dorsal genes such as gsc and chordin. Thus, the ventralizing activity of Cav-1 is mediated by its ability to induce ventral genes and inhibit dorsal genes. |
|
Cav-1 proteins from different species mediate conserved ventralizing activity. (A) Amino acid sequence analysis of human and zebrafish Cav-1 (GenBank # of human Cav-1α and -1β: NP_001744 and ABW22626; GenBank # of zebrafish Cav-1α and -1β: NP_997816 and NP_001019333). Caveolin scaffolding domain was shown in a red box. (B) Overexpression of human cav-1 ventralizes zebrafish embryos. Embryos at one-cell stage were injected with 300 pg human cav-1α or -1β mRNA. Total ventralized embryos (T) were divided into three groups: mild (M), intermediate (I) and severe (S). (C) Expression patterns of dorsoventral genes in embryos injected with 300 pg human cav-1α mRNA, 300 pg human cav-1β mRNA, 150 pg human cav-1α mRNA plus 10 ng zebrafish cav-1α-MO, or 150 pg human cav-1β mRNA plus 10 ng zebrafish cav-1β-MO. Developmental stages are shown at the bottom. Animal pole views of embryos at shield stage with the dorsal toward the right for chordin and eve1. Dorsal views of embryos at shield stage with animal pole toward the top for gsc. Lateral views of embryos at shield stage with the dorsal toward the right for bmp2b. Lateral views of embryos at 24 hpf with anterior to the left for gata1. (D–E)Percentage of embryos with decreased or increased expression of marker genes in (C). Total numbers of embryos examined are indicated at the bottom. (F) Co-injection of 150 pg human cav-1α mRNA into embryos at one-cell stage blocked activation of the Top-flash luciferase reporter by injection of 150 pg β-cat mRNA, but not that by injection of 150 pg β-cat-M mRNA. **indicates p < 0.01 vs. embryos injected with Top-flash plus β-cat mRNA. |
|
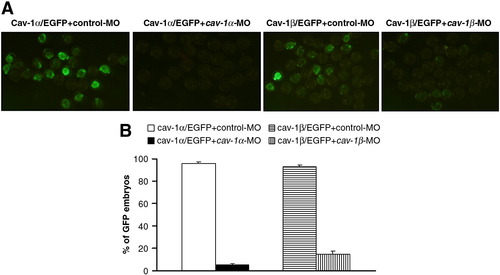
Efficiency of cav-1 knockdown by cav-1-MOs. (A) Embryos at one-cell stage were injected with 200 pg Cav-1α/EGFP plus 10 ng control-MO, 200 pg Cav-1α/EGFP plus 10 ng cav-1α-MO, 200 pg Cav-1β/EGFP plus 10 ng control-MO, or 200 pg Cav-1β/EGFP plus 10 ng cav-1β-MO. (B) Percentage of GFP-expressing embryos for each group. Data represent mean ± S.D. from three independent experiments. |
|
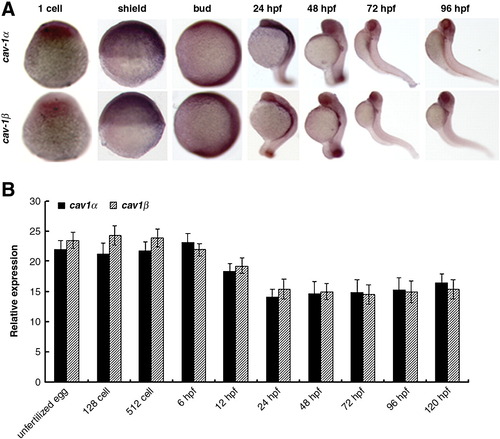
Spatiotemporal expression of cav-1 during embryogenesis. (A) Whole-mount in situ hybridization was performed with antisense RNA probes in embryos at indicated stages. 1-cell and shield stages, lateral view with the animal pole to the top; Bud stage to 96 hpf, lateral view with the anterior to the top. (B) Relative expression of Cav-1α or -1β during embryogenesis. Embryos at indicated stages were collected for total RNA isolation and subjected to real-time PCR analysis. Expression of 18s ribosomal RNA was used as an internal control. Values are given as mean ± standard deviation, n = 3. |
Reprinted from Developmental Biology, 344(1), Mo, S., Wang, L., Li, Q., Li, J., Li, Y., Thannickal, V.J., and Cui. Z., Caveolin-1 regulates dorsoventral patterning through direct interaction with beta-catenin in zebrafish, 210-223, Copyright (2010) with permission from Elsevier. Full text @ Dev. Biol.