- Title
-
A role for chemokine signaling in neural crest cell migration and craniofacial development
- Authors
- Olesnicky Killian, E.C., Birkholz, D.A., and Artinger, K.B.
- Source
- Full text @ Dev. Biol.
|
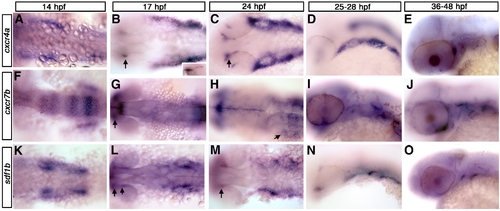
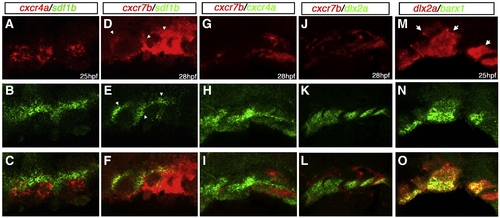
cxcr4a, cxcr7b, and sdf1b expression in the zebrafish embryo. Dorsal views (A, B, C, F, G, H, K, L, M), lateral views (D, E, I, J, N, O) and anterior to the left. (A–E) cxcr4a expression. cxcr4a is observed in NCCs migrating to the arches (A). At 17 hpf (B) and 24 hpf (C) is expressed in the optic stalk (B, C; arrows) and pharyngeal arch region but is excluded from the central arch, likely corresponding to the mesodermal core. By 25 hpf, cxcr4a expression condenses into discreet arches (D). At 36 hpf, cxcr4a is expressed in the optic stalk and throughout the arches (E). (F–J) cxcr7b expression. From 14–17 hpf, cxcr7b is expressed in the midbrain, otic placode and in rhombomeres 3, 5 and 6 and, at 17 hpf, in the optic stalk (F, G; arrow in G to optic stalk). By 24 hpf, cxcr7b is expressed in the ventral region of the posterior pharyngeal arches (H, arrow). From 28–48 hpf, cxcr7b expression is seen throughout the pharyngeal arches (I, J). (K–O) sdf1b expression. At 14–17 hpf, sdf1b is expressed in the region of the migrating CNCCs, within the optic stalk and dorsal rim of eye (K, L; arrows in L and M). By 24 hpf (M) sdf1b is expressed in the typical scalloped pattern of the pharyngeal endoderm. At 25 hpf, sdf1b is expressed in endodermal pouches 1–3 (N) and pouches 1 and 2 at 36 hpf (O). |
|
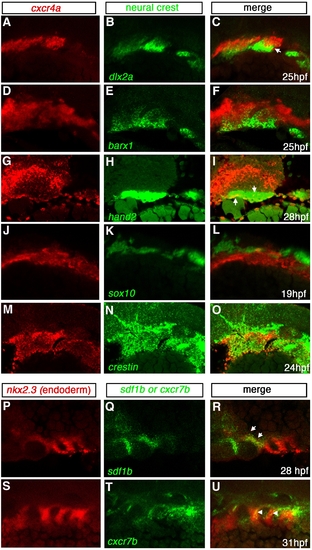
Expression of cxcr4a, cxcr7b and sdf1b in the developing pharyngeal arches. Lateral views, anterior to the left. Single channel (A, B, D, E, G, H, J, K, M, N, P, Q, S, T) and merged (C, F, I, L, O, R, U) images of confocal micrographs of double fluorescent in situ hybridization of cxcr4a, cxcr7, and sdf1b along with tissue specific marker expression in the pharyngeal arch region. Neural crest markers are indicated in green with cxcr4a (red) in A–O, and endodermal marker nkx2.3 (red) with sdf1b or cxcr7b (green) in P–U. (A–C) CNCC marker dlx2a expression (green) overlaps cxcr4a (red) in a subset of CNCCs (yellow, arrows) at 25 hpf. (D–F) barx1 is coexpressed in a subset of cxcr4a cells at 25 hpf. (G–I) At 28 hpf, hand2 is expressed in the ventral-most domain of arch CNCCs and partially overlaps with cxcr4a in this domain (yellow, arrows). (J–L) cxcr4a (red) is mostly excluded from sox10 nonectomesenchymal expression at 19 hpf (green). (M–O) The pan CNCC marker crestin is expressed in a broad domain in the pharyngeal arch region, where cxcr4a is also expressed. cxcr4a is excluded from the dorsal crestin domain that corresponds to nonectomesenchymal NCCs. (P–R) sdf1b (green) and the endodermal marker nkx2.3 (red) overlap in pouch 2 (arrows) at 28 hpf. (S–U) cxcr7b (green) is coexpressed with nkx2.3 (red) throughout pharyngeal arch endoderm at 31 hpf (yellow; arrowheads). |
|
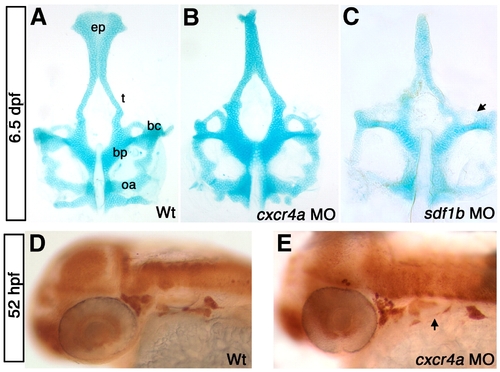
cxcr4a morphants have neurocranium defects. Dorsal views, anterior to the top (A–C) and lateral views, anterior to the left (D, E). (A) wild type (WT) neurocranium at 6.5 dpf. (B) cxcr4a morphants lack an ethmoid plate (ep) and have fused trabeculae (t). (C) sdf1b morphants phenocopy cxcr4a morphants, and have more severe anterior basicapsular commissure defects (arrow). (D, E) Wildtype (D) and cxcr4a morphant embryo (E) labeled with anti HuC/D 52 hpf to indicate the cranial ganglia. Morphant embryos have a reduction in cranial ganglia IX and X (arrow). bc, basicapsular commissure; bp, basil plate; oa, occipital arch; ep, ethmoid plate; t, trabeculae. |
|
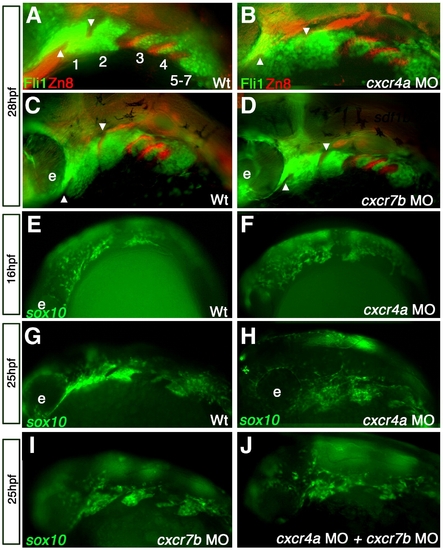
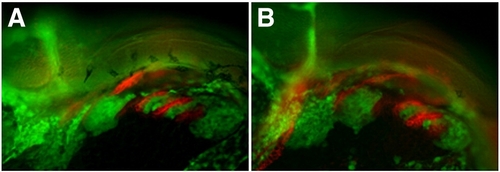
cxcr4a morphants have aberrant CNCC migration and arch condensation. Lateral views, anterior to the left. (A–D) tg{fli1::eGFP} (green) embryo marking postmigratory CNCCs double labeled with the endoderm marker Zn-8 (red). (E–J) tg{sox10::eGFP}(green) labeling neural crest migration and condensation into the pharyngeal arch region. (A, B) cxcr4a morphants (B) show loose arch structure, as compared to wild type embryos. (A) Arrowheads mark endodermal pouch 1 and stomodeum in A–D as a reference for compaction. In addition, endodermal specification and morphogenesis is unaffected as seen by Zn8 expression (red). (C, D) cxcr7b MO injected embryos (D) have normal CNCC (green) compaction in the pharyngeal arch and endodermal expression of Zn8 (red) as compared to wildtype at 28 hpf (C). (E, G) Wildtype tg{sox10::eGFP} embryos display normal onset of CNCC migration from the neural tube toward the ventral arch region, migrate normally mid-migration at 16 hpf (E) and are properly organized post-migration at 25 hpf (G). Early to mid CNCC migration is unaffected in cxcr4a morphants at 16 hpf, as CNCCs are migrating ventrally from the neural tube (F) but show loose organization of CNCC in the anterior arches at 25 hpf (H). cxcr7b morphant embryos have normal arch morphology (I) and double cxcr4a and cxcr7b morphants (J) show a similar disorganized arch phenotype as cxcr4a single morphants. |
|
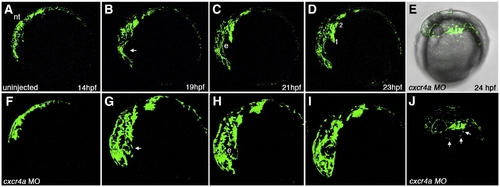
Time lapse imaging of aberrant CNCC migration in cxcr4a morphant embryos. Lateral views, anterior to the left. Stills taken from live time lapse imaging from 14–23 hpf. Wildtype CNCC migration from 14 to 23 hpf (A–D). Wildtype CNCCs avoid the eye during early CNCC migration (B, arrow). cxcr4a morphant CNCCs do not avoid the eye during early phases of migration (F–I; B, arrow). (E, J) Brightfield and fluorescent (E) and fluorescence only (J) images of cxcr4a morphant CNCCs migrating ectopically over the yolk. e, eye. Numbers denote arches. |
|
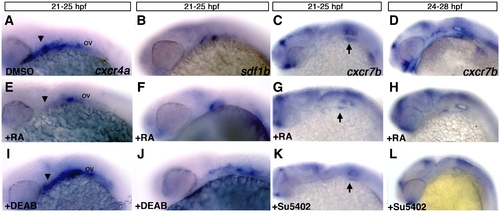
Retinoic acid signaling is upstream of cxcr4a and sdf1b. Lateral views, anterior to the left of 25 hpf (A–C, E–G, I–K) and 28 hpf embryos (D, H, L). (A–D) Control embryos treated with DMSO between 21 and 25 hpf show normal expression of cxcr4a (A), sdf1b (B), and from 21–25 hpf and 24–28 hpf showing normal cxcr7b expression (C, D respectively). Embryos treated with 100 μM RA from 21–25 hpf show reduced expression of cxcr4a (E as compared to A; arrowheads), a mild reduction in pouch 2 of sdf1b (F) and mild reduction of cxcr7b (G) below the otic vesicle (arrows point to otic vesicle region in C, G, K). Treatment from 24–28 hpf (H) causes a reduction of cxcr7b expression in the endoderm but not the forebrain as compared to wildtype embryos. Embryos treated with the 50 μM RA synthesis inhibitor DEAB show an expansion of cxcr4a (I) and sdf1b (J) expression posterior to the otic vesicle (ov) as compared to DMSO treated embryos in (A, B). Embryos treated with the Fgf inhibitor Su5402 from 21–25 hpf (K) and 24–28 hpf (L) have reduced expression of cxcr7b in the otic region (K, L) and within the pharyngeal endoderm (L) as compared to DMSO treated controls (C, D). |
|
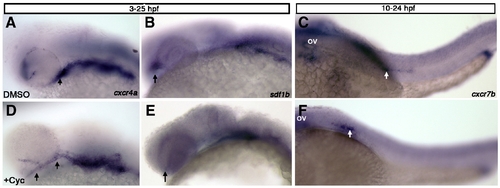
Hedgehog signaling regulates expression of sdf1b within the optic stalk. Lateral views, anterior to the left observed at 25 hpf (A, B, D, E) and 28 hpf (C, F). (A, B) Embryos treated with DMSO or cyclopamine (D, E) from 3 hpf–25 hpf show mispositioned cxcr4a expression within the optic stalk and less expression in the first arch of cyclopamine treated embryos (D; arrows), as compared to DMSO treated control embryos (A). sdf1b expression in the optic stalk is lost in cyclopamine treated animals (E; arrow) in comparison to DMSO treated controls (B; arrows point to optic stalk in B, E). (C, F) Embryos treated with cyclopamine from 10–24 hpf and observed at 28 hpf with cxcr7b expression show lateral line migration defects. The lateral line primordium in cyclopamine treated embryos (F) does not migrate posteriorly as compared to DMSO treated embryos (C; white arrows). ov, otic vesicle. |
|
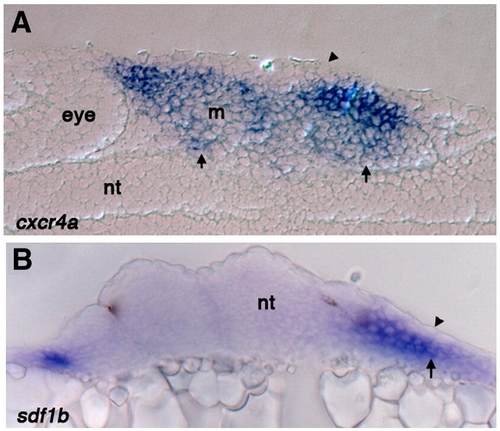
Sections through the pharyngeal arch region show cxcr4a and sdf1b expression is excluded from the ectoderm. (A) Sagittal section of 25 hpf wild type embryo stained for cxcr4a mRNA expression, arrow points to expression throughout the mesenchyme and arrowheads shows exclusion from the ectoderm. cxcr4a is also significantly reduced in the mesodermal core (B) Transverse section of 25 hpf wild type embryo stained for sdf1b expression. Expression is absent from the ectoderm (arrowhead) but present in the endoderm (arrow). Nt, neural tube; m, mesoderm. |
|
Double in situ hybridization of cxcr4a/cxcr7b/sdf1b in the developing pharyngeal arches. Lateral views, anterior to the left. Confocal micrographs of double fluorescent in situ hybridization of cxcr4a, cxcr7, sdf1b, dlx2a and barx1 in the pharyngeal arch region. (A–C) cxcr4a (red) is largely excluded from sdf1b (green) in the arches, but is slightly coexpressed with sdf1b at the endodermal border. (D–F) cxcr7b (red) is coexpressed with sdf1b (green) throughout the medial endoderm and anterior pharyngeal pouches at 28 hpf (arrowheads). (G–I) cxcr4a (green) is mostly excluded from cxcr7b (red) expressing cells, but does show some coexpression in the posterior pharyngeal region. (J–L) cxcr7b (red) is excluded from the dlx2a expressing CNCCs in the anterior arches but does show partial overlap with dlx2a expressing cells of the posterior pharyngeal arches. (M–O) barx1 is coexpressed with dlx2a in the ventral and medial portion of the pharyngeal arches but is reduced or absent in the dorsal arch (arrows). |
|
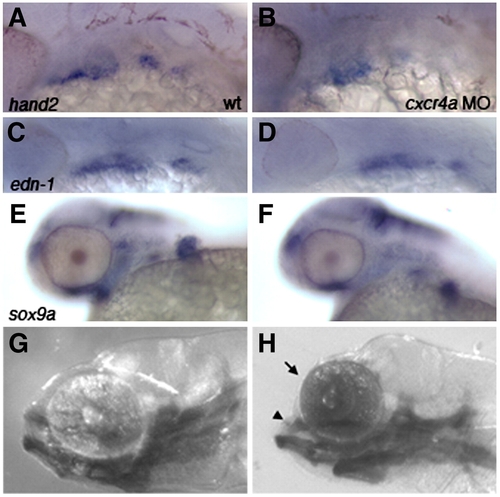
cxcr4a morphants have a “dolphin” phenotype and reduced arch marker expression. Lateral views, anterior to the left. Expression of hand2, edn-1 and sox9a in wildtype (A, C, E) and cxcr4a morphant embryos (B, D, F) shows that expression is reduced but not absent after cxcr4a knockdown. (G) Wildtype larva at 6.5 dpf. (H) cxcr4a morphant larva at 6.5 dpf has a reduced forebrain and “dolphin” like phenotype (arrow and arrowhead). |
|
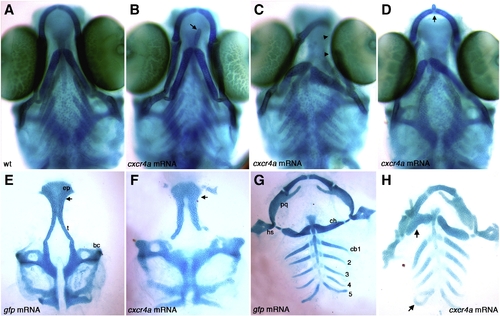
Overexpression of cxcr4a causes ectopic cartilage formation. Whole-mount (A–D) and dissected (E–H) alcian blue stained cartilage at 6.5 dpf, anterior to the top. Wildtype (A) and gfp mRNA injected controls (E,G) and examples of cxcr4a mRNA injected embryos (B–D,F,H). Injection of 100 pg cxcr4a mRNA results in small ectopic cartilages often located near the Meckel′s and ceratohyal cartilages (B, D; arrow). We also often observe unilateral loss of Meckel′s cartilages and abnormal palatoquadrate elements (C; arrowheads). Injection of 250 pg cxcr4a mRNA results in abnormal hyosymplectic elements (hs), fusions of the Meckel′s and ceratohyal cartilages, as well as fusions of the posterior ceratobranchial elements (H, arrow) as compared to control embryos injected with gfp mRNA (G). Injection of 100 pg of cxcr4a mRNA causes ectopic cartilages and neurocranium defects with a reduction of the trabeculae (t) and a cleft within the ethmoid plate (ep; F; arrow) as compared to control embryos injected with gfp mRNA (E; arrow at same location). bc, basicapsular commissure; cb1–5, ceratobranchial cartilage 1–5; ch, ceratohyal; hs, hyosymplectic; pq, palatoquadrate; ep, ethmoid plate; t, trabeculae. |
|
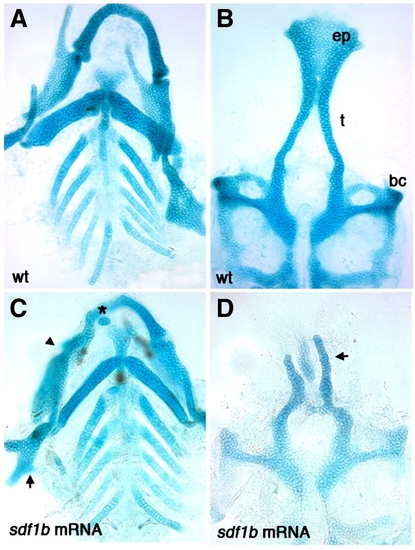
sdf1b overexpression causes defects in the larval craniofacial skeleton. Dissected alcian blue stained cartilage at 6.5 dpf, anterior to the top. Wildtype control larva (A, B) and sdf1b mRNA injected larvae (C, D). Injection of 250 pg sdf1b mRNA results in small ectopic cartilages (C asterisk, arrow, D), abnormal hyosymplectic elements (C, arrow), fusions of the Meckel′s and palatoquadrate cartilages (C, arrowhead) as compared to control larvae (A). sdf1b mRNA overexpression causes ectopic cartilages and neurocranium defects with a reduction of the trabeculae, a clefted palate and loss of the ethmoid plate (D), as compared to control larvae (B). |
|
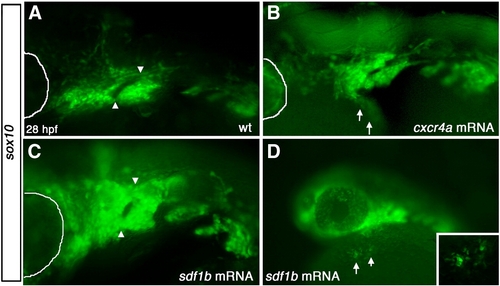
Overexpression of sdf1b and cxcr4a results in CNCC defects. Lateral views, anterior to the left of 28 hpf embryos (A,B). (A) WT tg{sox10::eGFP} embryos show normal pharyngeal arch patterning, while embryos injected with cxcr4a mRNA (B) show pharyngeal arch defects, with much of arch 1 missing and ectopic CNCCs ventro-lateral to the arch region along the yolk (arrows). Embryos injected with sdf1b mRNA (C) show similar arch defects to cxcr4a mRNA overexpression with fusion of arch 1 and 2. Lower (10x) magnification of embryo injected with sdf1b mRNA showing ectopic CNCCs along the yolk ventral to the eye and pharyngeal arches (D, arrows). Inset shows 20x magnification of ectopic CNCCs along yolk. eye is outlined with white line in panels A–C. |
|
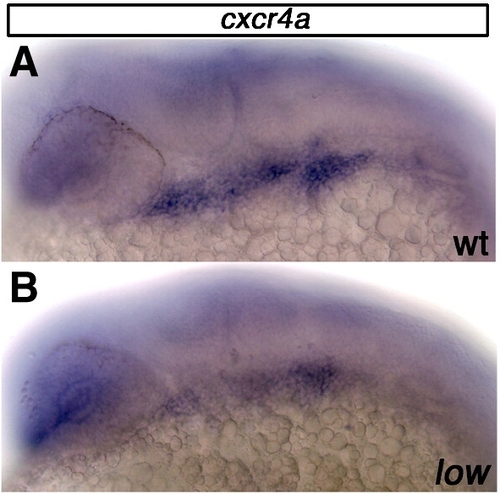
cxcr4a expression is reduced in lockjaw mutant embryos at 25 hpf. (A) Wildtype embryo stained for cxcr4a mRNA expression 25 hpf. (B) lockjaw mutant embryos have reduced cxcr4a expression within the pharyngeal arches at 25 hpf. |
|
Endodermal development following SU5402 treatment from 24–28 hpf. Lateral views, anterior to the left. (A,B) tg{fli1::eGFP} (green) embryo marking postmigratory CNCCs double labeled with the endoderm marker Zn-8 (red). (A) Control embryo treated from 24–28 hpf with DMSO shows normal arch and pharyngeal pouch development. (B) Embryo treated with SU5402 from 24–28 hpf shows reduced arches 3 and 4, as well as broad and less elongated posterior pharyngeal pouches as compared to DMSO treated control sibling. |
|
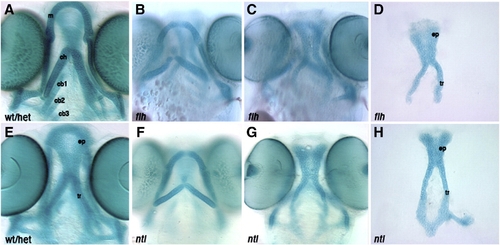
Larval head skeletons of floating head and notail mutants do not phenocopy cxcr4a or sdf1b morphants. Whole-mount (A–C, E–G) and dissected (D,H) alcian blue stained cartilage at 4.5 dpf, anterior to the top. Ventral cartilages of a wildtype or heterozygous larva (A). floating head (flt) (B) and notail (ntl) mutant larvae (F) show normal anterior ventral cartilage development, but are missing posterior ceratobranchials. floating head (C) and notail (G) mutant larvae form an ethmoid plate and trabeculae, but are missing the posterior neurocranium as compared to control neurocranium (E). Dissected neurocranium of floating head (D) and notail mutant larvae (H) showing a normal ethmoid plate. wt/het, wildtype or heterozygous; m, Meckel′s cartilage; ch, ceratohyal; cb1, 2, 3, ceratobranchial 1, 2, 3; ep, ethmoid plate; tr, tribeculae. |
Reprinted from Developmental Biology, 333(1), Olesnicky Killian, E.C., Birkholz, D.A., and Artinger, K.B., A role for chemokine signaling in neural crest cell migration and craniofacial development, 161-172, Copyright (2009) with permission from Elsevier. Full text @ Dev. Biol.