- Title
-
Developmental patterning of the cardiac atrioventricular canal by Notch and Hairy-related transcription factors
- Authors
- Rutenberg, J.B., Fischer, A., Jia, H., Gessler, M., Zhong, T.P., and Mercola, M.
- Source
- Full text @ Development
|
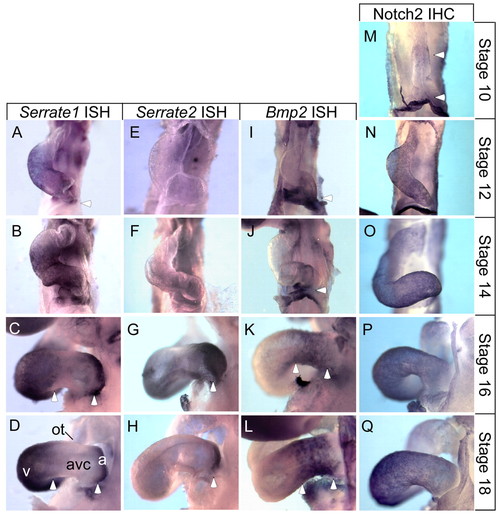
Serrate1 and Serrate2 gene expression patterns correspond to the borders of the AVC and indicate prospective atrial and ventricular chamber myocardium. (A-D) Stage 12-18 embryos were hybridized with a Serrate1 antisense RNA probe. (A) Stage 12 embryos exhibit widespread Serrate1 expression throughout the posterior heart with the anterior boundary at the outflow tract. The posterior border stops at the presumptive atrium and sinus venosus, but may include primitive AVC tissue (white arrowhead). (B,C) Between stages 14 and 16 expression becomes restricted to the common ventricle and atria with boundaries at the AVC (white arrowheads, C). (D) At stage 18, expression is strong in the ventricles and atria, forming distinct boundaries at both the anterior and posterior AVC border (white arrowheads). (E-H) Stage 12-18 embryos were hybridized with a Serrate2 antisense RNA probe. (E,F) At early stages, 12 and 14, Serrate2 is not detected in the heart. (G) Serrate2 expression becomes apparent by stage 16 at the posterior-most region of the heart, corresponding to the distal atrium and sinus venosus (white arrowhead). (H) By stage 18, strong Serrate2 expression is observed in the atria and sinus venosus, overlapping that of Serrate1 (white arrowhead). (I-L) Stage 12-18 embryos were hybridized with the AVC marker Bmp2 for comparison with the Notch ligands. (I,J) Bmp2 is expressed in the posterior-most region of the heart, including the primitive AVC, atria and sinus venosus. The most anterior Bmp2 expression appears to correspond to the posterior border of Serrate1 expression (compare arrowheads in A,B and I,J). (J,K) Between stages 14 and 16, Bmp2 expression is restricted to the AVC. Again, Bmp2 expression appears complementary to that of Serrate1 in the primitive ventricle, and to both Serrate1 and Serrate2 in the atria (compare arrowheads in K with C,G). (L) Bmp2 mRNA persists in the AVC. (M-Q) Immunostaining against Notch2. (M) Notch2 is first detected in the linear heart tube in small patches in the ventral primitive ventricle and at the junction of the two horns of the primitive atria (white arrowheads). (N-Q) By Stage 12, Notch2 is present throughout the myocardium, where it remains at all stages examined. a, atrial region; avc, atrioventricular canal; ot, outflow tract; v, ventricular region. |
|
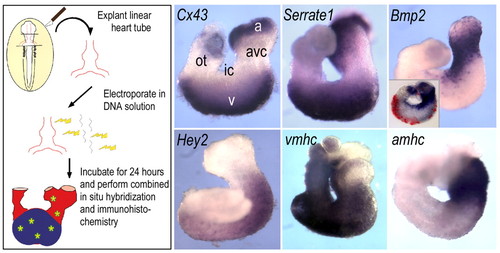
Plasmid electroporation of linear heart tubes. Stage 12 hearts were dissected, electroporated with RSV promoter cDNA constructs to direct production of epitope-tagged Notch pathway modulators or GFP (see Materials and methods). During the subsequent 24-hour culture period, the electroporated hearts looped and developed appropriate expression patterns of region-specific markers, including Cx43, vmhc and amhc in chamber myocardium, as well as Bmp2 in the AVC. In addition, Serrate1 and Hey2 showed characteristic expression patterns. Inset in Bmp2 panel shows Cy3 immunofluorescent detection of the Myc-epitope tag-labelled product of transfected eGFP gene. a, atrial region; avc, atrioventricular canal; ic, inner curvature; ot, outflow tract; v, ventricular region. |
|
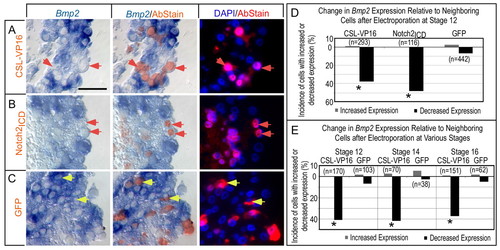
Notch pathway agonists inhibit Bmp2 expression cell autonomously in the AVC and IC. (A-C) Histological sections demonstrating that misactivation of Notch signaling suppresses Bmp2. Immunohistochemical staining (red fluorescence) detected epitopes on expressed proteins from the electroporated cDNAs: CSL-VP16 (A), Notch2ICD (B) or GFP control (C). Brightfield images on the left show in situ hybridization (blue). Cy3 (red) and DAPI (blue) fluorescent images are shown on right. The center panels show brightfield images merged with the fluorescent Cy3 (red) staining of the epitope-tagged proteins. Red arrows indicate examples of cells with reduced Bmp2 relative to neighbors, whereas yellow arrows indicate examples of unchanged cells. Scale bar in A: 50 μM. (D,E) Quantification of cells scored for altered gene expression relative to neighboring cells after electroporation with the indicated cDNAs, scored from histological sections as in A-C. All sample identifiers were encoded prior to histological embedding so that tissue processing and scoring was performed blind (see Materials and methods). Data plotted are cumulative over greater than three trials. Note that Notch pathway activation suppresses Bmp2 expression (D) and that Bmp2 remains responsive throughout the period of heart looping (HH stages 12-16; E). Asterisks indicate statistical significance of the difference between the experimental modulation of Notch pathway and the control GFP (P<0.05, χ2 test). |
|
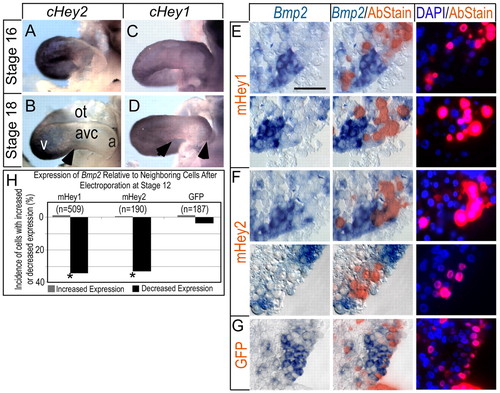
Ectopic Hey1 and Hey2 suppress Bmp2 cell autonomously in the AVC and IC. (A-D) Examples of endogenous Hey2 and Hey1 mRNA expression in stage 16 and 18 embryos. Arrowheads denote the borders of mRNA expression. (A,B) Hey2 expression is restricted to the developing ventricles at all stages examined, similar to patterns reported for the mouse. (C) Low levels of Hey1 are detected throughout the myocardium of stage 16 chick hearts. (D) By stage 18, Hey1 is detected in atrial and ventricular myocardium, but not AVC, as for expression of Serrate1 (see Fig. 1). Arrowheads indicate the borders of Hey gene expression with the AVC. (E-G) Bmp2 suppression by exogenously provided Myc-tagged mouse Hey1 and Flag-tagged mouse Hey2. Immunohistochemical staining (red fluorescence) detected epitopes on expressed proteins from the electroporated cDNAs, as in Fig. 3: Hey1 (E), Hey2 (F) or GFP control (G). Scale bar in E: 50 μM. (H) Quantification was performed as for Fig. 3. Asterisk indicates statistical significance of a difference between the effect of Hey proteins and GFP control (P<0.05, χ2 test). |
|
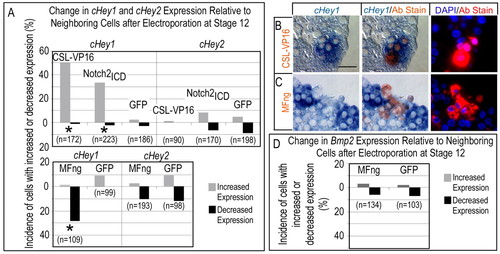
Differential effects of Notch on Hey1, Hey2 and Bmp2. Stage 12 hearts were electroporated to misexpress modulators of the Notch pathway signaling, cultured for 24 hours and analyzed for effects on endogenous gene expression, as described in Fig. 3. (A) Activation of the Notch pathway with Myc-tagged CSL-VP16 or Notch2icd increased the number of cells that expressed endogenous Hey1, relative to neighboring cells, and attenuation of Notch with HA-tagged Mfng caused a decrease in Hey1-positive cells. By contrast, Notch pathway agonists and antagonists did not affect Hey2 expression. (B,C) Histological sections demonstrating that misactivation of the Notch pathway (B) or inhibition of endogenous Notch responsiveness by Mfng (C) elicited reciprocal and cell-autonomous effects on endogenous Hey1. Scale bar: 25 μm. (D) Attenuation of Notch signaling with Mfng did not cause a cell-autonomous effect on the expression of Bmp2, consistent with the observations that Hey2 is not regulated directly by Notch (A) yet suppresses Bmp2 (see Fig. 4). Asterisks in A,D indicate statistical significance of the difference between the Notch pathway modulators and the GFP control (P<0.05, χ2 test). |
|
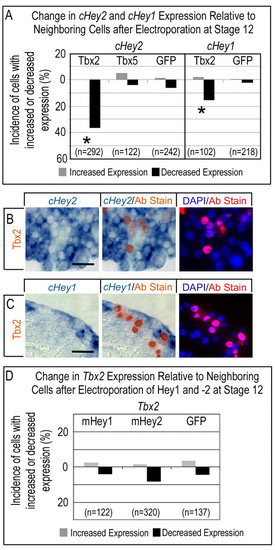
Tbx2 represses Hey1 and Hey2 cell autonomously. (A) Misexpression of Myc-tagged Tbx2 in stage 12 hearts caused a cell-autonomous reduction in the incidence of Hey1- and Hey2-positive cells. Tbx5, by contrast, had no effect. Asterisks indicate statistical significance of the difference between electroporated Tbx2 and the GFP control (P<0.05, χ2 test). (B,C) Histological sections demonstrating the cell-autonomous suppression of endogenous Hey1 (B) and Hey2 (C) by electroporated Tbx2. (D) Flag-tagged mouse Hey1 and mouse Hey2 did not alter Tbx2 expression. Brightfield, merged and fluorescent images are as in Fig. 3. Scale bar: 50 μm. |
|
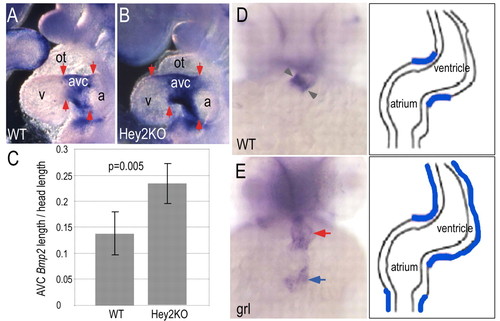
Expansion of the cardiac Bmp expression domain in the absence of Hey2 homologs in mouse and zebrafish embryos. (A,B) Bmp2 expression marks the AVC and IC of E9.5-E10.0 mouse wild-type (WT) embryos (A), and this region is expanded in homozygous Hey2 mutant siblings (B). Arrows indicate the borders of Bmp2 expression. (C) Measurements of the atrioventricular length of the Bmp2 expression domain relative to head length revealed a 1.7-fold expansion in mutants (n=5) relative to wild type (n=5; P<0.005, Student's t-test), compiled from two litters. Error bars correspond to standard deviation. (D,E) Bmp4 marked the developing AVC (gray arrowheads) in 48 hpf wild-type zebrafish embryos (D), whereas gridlock (grl) mutant embryos (E), which lack the Hey2 homolog, showed diffusely distributed Bmp4 in the AVC and ventricular regions (red arrow), and strong ectopic expression in the inflow tract (blue arrow), as diagrammed in schematics to the right. EXPRESSION / LABELING:
|